|
ATCC
204305 a fumigatus strain 204305 A Fumigatus Strain, supplied by ATCC, used in various techniques. Bioz Stars score: 98/100, based on 1 PubMed citations. ZERO BIAS - scores, article reviews, protocol conditions and more https://www.bioz.com/result/204305 a fumigatus strain/product/ATCC Average 98 stars, based on 1 article reviews
204305 a fumigatus strain - by Bioz Stars,
2026-03
98/100 stars
|
Buy from Supplier |
|
ATCC
reference strain e coli atcc 25922  Reference Strain E Coli Atcc 25922, supplied by ATCC, used in various techniques. Bioz Stars score: 99/100, based on 1 PubMed citations. ZERO BIAS - scores, article reviews, protocol conditions and more https://www.bioz.com/result/reference strain e coli atcc 25922/product/ATCC Average 99 stars, based on 1 article reviews
reference strain e coli atcc 25922 - by Bioz Stars,
2026-03
99/100 stars
|
Buy from Supplier |
|
Addgene inc
red glifon 300 3000  Red Glifon 300 3000, supplied by Addgene inc, used in various techniques. Bioz Stars score: 93/100, based on 1 PubMed citations. ZERO BIAS - scores, article reviews, protocol conditions and more https://www.bioz.com/result/red glifon 300 3000/product/Addgene inc Average 93 stars, based on 1 article reviews
red glifon 300 3000 - by Bioz Stars,
2026-03
93/100 stars
|
Buy from Supplier |
|
ATCC
monkeypox virus mpxv iib strain mpxv france irba2211i 2022  Monkeypox Virus Mpxv Iib Strain Mpxv France Irba2211i 2022, supplied by ATCC, used in various techniques. Bioz Stars score: 94/100, based on 1 PubMed citations. ZERO BIAS - scores, article reviews, protocol conditions and more https://www.bioz.com/result/monkeypox virus mpxv iib strain mpxv france irba2211i 2022/product/ATCC Average 94 stars, based on 1 article reviews
monkeypox virus mpxv iib strain mpxv france irba2211i 2022 - by Bioz Stars,
2026-03
94/100 stars
|
Buy from Supplier |
|
ATCC
e coli type strain atcc 11775 in vitro  E Coli Type Strain Atcc 11775 In Vitro, supplied by ATCC, used in various techniques. Bioz Stars score: 99/100, based on 1 PubMed citations. ZERO BIAS - scores, article reviews, protocol conditions and more https://www.bioz.com/result/e coli type strain atcc 11775 in vitro/product/ATCC Average 99 stars, based on 1 article reviews
e coli type strain atcc 11775 in vitro - by Bioz Stars,
2026-03
99/100 stars
|
Buy from Supplier |
|
GE Healthcare
e coli bl21 strain de3  E Coli Bl21 Strain De3, supplied by GE Healthcare, used in various techniques. Bioz Stars score: 94/100, based on 1 PubMed citations. ZERO BIAS - scores, article reviews, protocol conditions and more https://www.bioz.com/result/e coli bl21 strain de3/product/GE Healthcare Average 94 stars, based on 1 article reviews
e coli bl21 strain de3 - by Bioz Stars,
2026-03
94/100 stars
|
Buy from Supplier |
|
ATCC
dmso  Dmso, supplied by ATCC, used in various techniques. Bioz Stars score: 96/100, based on 1 PubMed citations. ZERO BIAS - scores, article reviews, protocol conditions and more https://www.bioz.com/result/dmso/product/ATCC Average 96 stars, based on 1 article reviews
dmso - by Bioz Stars,
2026-03
96/100 stars
|
Buy from Supplier |
|
ATCC
vitro test ozone gas standard strains s aureus  Vitro Test Ozone Gas Standard Strains S Aureus, supplied by ATCC, used in various techniques. Bioz Stars score: 99/100, based on 1 PubMed citations. ZERO BIAS - scores, article reviews, protocol conditions and more https://www.bioz.com/result/vitro test ozone gas standard strains s aureus/product/ATCC Average 99 stars, based on 1 article reviews
vitro test ozone gas standard strains s aureus - by Bioz Stars,
2026-03
99/100 stars
|
Buy from Supplier |
|
ATCC
helicobacter pylori strain atcc 43504  Helicobacter Pylori Strain Atcc 43504, supplied by ATCC, used in various techniques. Bioz Stars score: 99/100, based on 1 PubMed citations. ZERO BIAS - scores, article reviews, protocol conditions and more https://www.bioz.com/result/helicobacter pylori strain atcc 43504/product/ATCC Average 99 stars, based on 1 article reviews
helicobacter pylori strain atcc 43504 - by Bioz Stars,
2026-03
99/100 stars
|
Buy from Supplier |
|
Vector Laboratories
biotinylated dolichos biflorus agglutinin dba  Biotinylated Dolichos Biflorus Agglutinin Dba, supplied by Vector Laboratories, used in various techniques. Bioz Stars score: 94/100, based on 1 PubMed citations. ZERO BIAS - scores, article reviews, protocol conditions and more https://www.bioz.com/result/biotinylated dolichos biflorus agglutinin dba/product/Vector Laboratories Average 94 stars, based on 1 article reviews
biotinylated dolichos biflorus agglutinin dba - by Bioz Stars,
2026-03
94/100 stars
|
Buy from Supplier |
|
ATCC
genomic dna  Genomic Dna, supplied by ATCC, used in various techniques. Bioz Stars score: 99/100, based on 1 PubMed citations. ZERO BIAS - scores, article reviews, protocol conditions and more https://www.bioz.com/result/genomic dna/product/ATCC Average 99 stars, based on 1 article reviews
genomic dna - by Bioz Stars,
2026-03
99/100 stars
|
Buy from Supplier |
Image Search Results
 Fig. S2 A . In dark gray , P. aeruginosa SG17M specific tLST genes; light gray , additional tLST genes present in other tLST classes. C , molecular weight determination of PaFtsH1 and PaFtsH2 purified from Escherichia coli . Purified PaFtsH1 and PaFtsH2 were run on an analytical Superdex 200 3.2/3.0 gel filtration column ( left ) and separated by a 4 to 20% gradient SDS-PAGE in a two-fold dilution series (insets). Molecular weights of PaFtsH enzymes were consistent with hexamer formation plus ∼3.5 NP-40 micelles per PaFtsH1-6xHis hexamer (PaFtsH1 6 -6xHis: 743,373 Da; one NP-40 micelle: ∼90 Da) and ∼2.7 NP-40 micelles per PaFtsH2-6xHis hexamer (PaFtsH2-6xHis: 659,792 Da). Theoretical molecular weights of 6xHis-tagged PaFtsH monomers: PaFtsH1-6xHis: 70,886 Da; PaFtsH2-6xHis: 69,521 Da. D , hydrolysis of ATP (5 mM) by PaFtsH1 ( blue bars) or PaFtsH2 ( purple bars) (0.43 μM) in the presence or absence of the substrate β-casein (40 μM) at 40 °C. ATP hydrolysis rates by PaFtsH1 and PaFtsH2 increased in the presence of β-casein (two-tailed Mann-Whitney U test (∗ p < 0.05)). The rate of ATP hydrolysis by β-casein in the absence of enzyme was negligible (0.0032 ± 0.0004 ATP min −1 ). E , Forty micromolar β-casein was incubated at 40 °C with PaFtsH1 or PaFtsH2 (0.43 μM), and degradation kinetics were monitored by a 4 to 20% gradient SDS-PAGE. Data points are averages of three independent replicates ± SD ( bottom ). M: Molecular weight marker taken from the PaFtsH2 casein degradation gel. F , the N-terminal domain of phage λ repressor (λcI N ) with a C-terminal ssrA degron tag (λcI N -ssrA; 15 μM) was incubated at 40 °C with EcFtsH, PaFtsH1, or PaFtsH2 (3.04 μM). λcI N -ssrA PDB: 1LM . M: Molecular weight marker. G , SsrA degron-tagged Arc repressor-(6 x H)KNQHE (Arc-st11-ssrA; 15 μM) was incubated at 40 °C with EcFtsH, PaFtsH1, or PaFtsH2 (3.53 μM). Arc-st11-ssrA PDB: 1ARR . M: Molecular weight marker. D – G , all reactions used C-terminal 6xHis-tagged proteins. E - G , all reactions contained 5 mM ATP and a creatine kinase (CK)-based ATP regeneration system . DNTR, diverse N-terminal region; MC, membrane-cytoplasmic; MC 7+ , membrane-cytoplasmic linker plus seven C-terminal amino acids into the AAA+ ATPase module boundary; MW, Molecular weight; M, Molecular weight marker; mAU, milli-Absorbance Unit; MC 7+ , membrane-cytoplasmic plus next seven C-terminal amino acids; Peri, periplasmic domain; TM1, transmembrane helix 1; TM2, transmembrane helix 2. " width="100%" height="100%">
Fig. S2 A . In dark gray , P. aeruginosa SG17M specific tLST genes; light gray , additional tLST genes present in other tLST classes. C , molecular weight determination of PaFtsH1 and PaFtsH2 purified from Escherichia coli . Purified PaFtsH1 and PaFtsH2 were run on an analytical Superdex 200 3.2/3.0 gel filtration column ( left ) and separated by a 4 to 20% gradient SDS-PAGE in a two-fold dilution series (insets). Molecular weights of PaFtsH enzymes were consistent with hexamer formation plus ∼3.5 NP-40 micelles per PaFtsH1-6xHis hexamer (PaFtsH1 6 -6xHis: 743,373 Da; one NP-40 micelle: ∼90 Da) and ∼2.7 NP-40 micelles per PaFtsH2-6xHis hexamer (PaFtsH2-6xHis: 659,792 Da). Theoretical molecular weights of 6xHis-tagged PaFtsH monomers: PaFtsH1-6xHis: 70,886 Da; PaFtsH2-6xHis: 69,521 Da. D , hydrolysis of ATP (5 mM) by PaFtsH1 ( blue bars) or PaFtsH2 ( purple bars) (0.43 μM) in the presence or absence of the substrate β-casein (40 μM) at 40 °C. ATP hydrolysis rates by PaFtsH1 and PaFtsH2 increased in the presence of β-casein (two-tailed Mann-Whitney U test (∗ p < 0.05)). The rate of ATP hydrolysis by β-casein in the absence of enzyme was negligible (0.0032 ± 0.0004 ATP min −1 ). E , Forty micromolar β-casein was incubated at 40 °C with PaFtsH1 or PaFtsH2 (0.43 μM), and degradation kinetics were monitored by a 4 to 20% gradient SDS-PAGE. Data points are averages of three independent replicates ± SD ( bottom ). M: Molecular weight marker taken from the PaFtsH2 casein degradation gel. F , the N-terminal domain of phage λ repressor (λcI N ) with a C-terminal ssrA degron tag (λcI N -ssrA; 15 μM) was incubated at 40 °C with EcFtsH, PaFtsH1, or PaFtsH2 (3.04 μM). λcI N -ssrA PDB: 1LM . M: Molecular weight marker. G , SsrA degron-tagged Arc repressor-(6 x H)KNQHE (Arc-st11-ssrA; 15 μM) was incubated at 40 °C with EcFtsH, PaFtsH1, or PaFtsH2 (3.53 μM). Arc-st11-ssrA PDB: 1ARR . M: Molecular weight marker. D – G , all reactions used C-terminal 6xHis-tagged proteins. E - G , all reactions contained 5 mM ATP and a creatine kinase (CK)-based ATP regeneration system . DNTR, diverse N-terminal region; MC, membrane-cytoplasmic; MC 7+ , membrane-cytoplasmic linker plus seven C-terminal amino acids into the AAA+ ATPase module boundary; MW, Molecular weight; M, Molecular weight marker; mAU, milli-Absorbance Unit; MC 7+ , membrane-cytoplasmic plus next seven C-terminal amino acids; Peri, periplasmic domain; TM1, transmembrane helix 1; TM2, transmembrane helix 2. " width="100%" height="100%">
Journal: The Journal of Biological Chemistry
Article Title: The membrane-cytoplasmic linker defines activity of FtsH proteases in Pseudomonas aeruginosa clone C
doi: 10.1016/j.jbc.2023.105622
Figure Lengend Snippet: Purified hex americ PaFtsH1 and PaFtsH2 have distinct biochemical activities in vitro . A , schematic ( top ) and linear representation ( bottom ) of a prototypical hexameric FtsH enzyme in a bacterial cell with key regions labeled. The DNTR spans the N-terminus, TM1, periplasmic domain, TM2, MC linker, and the first seven amino acids in the AAA+ ATPase module (MC 7+ ). B , schematic of genetic loci of PaftsH genes in Pseudomonas aeruginosa SG17M clone C strain. PaFtsH1 is encoded on the core genome of SG17M ( blue ), and PaFtsH2 ( purple ) is encoded on a horizontally acquired genomic island termed the transmissible locus of stress tolerance (tLST) which is distinctive in SG17M clone C. Names of genes in the vicinity of ftsH1 and ftsH2 are labeled in
Article Snippet: For protein induction, EcFtsH-Myc-6xHis, PaFtsH1-6xHis, and PaFtsH2-6xHis variants were expressed from a pET-28a(+) plasmid in
Techniques: Purification, In Vitro, Labeling, Molecular Weight, Filtration, SDS Page, Two Tailed Test, MANN-WHITNEY, Incubation, Marker, Membrane
 Table S1 . Circled spots indicate peptides containing residues A132 and/or K135, which were previously shown to be important for in vivo Escherichia coli FtsH (EcFtsH) binding to RpoH (conserved RpoH residues from E. coli are A131 and K134; ( , )). B , amino acid sequence of PaRpoH ( top ). Residues in red indicate regions bound by 35 S-EcFtsH, 35 S-PaFtsH1, 35 S-PaFtsH2. Residues in blue correspond to A132 and K135 (see panel A legend). FtsH-bound sequences identified by peptide blotting are mapped on the predicted PaRpoH structure ( bottom ; ) and numbered as on the sequence above. C , colony sizes of wild-type P. aeruginosa SG17M, the SG17M Δ ftsH1 Δ ftsH2 double deletion and SG17M Δ ftsH1 Δ ftsH2 strains complemented with either an empty pJN105 expression vector (EV) or pJN105-derived expression of PaFtsH1 or PaFtsH2. Pa ftsH1 and Pa ftsH2 genes had been cloned into the expression vector pJN105 under the translational regulation of the Pa ftsH2 Shine Dalgarno (SD) sequence. “+” and “-” scoring system relates the qualitative degree of colony growth of a particular strain genotype relative to that of SG17M EV (++++; wild-type colony growth) or SG17M Δ ftsH1 Δ ftsH2 (-; very poor colony growth). For ( C and E ), numbers along the top refer to the serial dilution factor for plating cells. D , schematic representation of PaFtsH1 and PaFtsH2 hybrid protein constructs. Regions corresponding to PaFtsH1 and PaFtsH2 identity are depicted in gray and purple , respectively ( top ). Designation of protein variants and regions derived from PaFtsH1 and/or PaFtsH2 that make up the respective FtsH hybrid variants are indicated in the table ( bottom ). E , colony size of the P. aeruginosa SG17M Δ ftsH1 Δ ftsH2 double deletion strain complemented with Pa ftsH1 and Pa ftsH2 hybrid constructs as introduced in panel D . “+” and “-” scoring system is the same as in ( C ). DNTR, Diverse N-terminal region; MC 7+ , membrane-cytoplasmic linker plus seven C-terminal amino acids into the AAA+ ATPase module boundary; TM1, Transmembrane helix 1; TM2, Transmembrane helix 2. " width="100%" height="100%">
Table S1 . Circled spots indicate peptides containing residues A132 and/or K135, which were previously shown to be important for in vivo Escherichia coli FtsH (EcFtsH) binding to RpoH (conserved RpoH residues from E. coli are A131 and K134; ( , )). B , amino acid sequence of PaRpoH ( top ). Residues in red indicate regions bound by 35 S-EcFtsH, 35 S-PaFtsH1, 35 S-PaFtsH2. Residues in blue correspond to A132 and K135 (see panel A legend). FtsH-bound sequences identified by peptide blotting are mapped on the predicted PaRpoH structure ( bottom ; ) and numbered as on the sequence above. C , colony sizes of wild-type P. aeruginosa SG17M, the SG17M Δ ftsH1 Δ ftsH2 double deletion and SG17M Δ ftsH1 Δ ftsH2 strains complemented with either an empty pJN105 expression vector (EV) or pJN105-derived expression of PaFtsH1 or PaFtsH2. Pa ftsH1 and Pa ftsH2 genes had been cloned into the expression vector pJN105 under the translational regulation of the Pa ftsH2 Shine Dalgarno (SD) sequence. “+” and “-” scoring system relates the qualitative degree of colony growth of a particular strain genotype relative to that of SG17M EV (++++; wild-type colony growth) or SG17M Δ ftsH1 Δ ftsH2 (-; very poor colony growth). For ( C and E ), numbers along the top refer to the serial dilution factor for plating cells. D , schematic representation of PaFtsH1 and PaFtsH2 hybrid protein constructs. Regions corresponding to PaFtsH1 and PaFtsH2 identity are depicted in gray and purple , respectively ( top ). Designation of protein variants and regions derived from PaFtsH1 and/or PaFtsH2 that make up the respective FtsH hybrid variants are indicated in the table ( bottom ). E , colony size of the P. aeruginosa SG17M Δ ftsH1 Δ ftsH2 double deletion strain complemented with Pa ftsH1 and Pa ftsH2 hybrid constructs as introduced in panel D . “+” and “-” scoring system is the same as in ( C ). DNTR, Diverse N-terminal region; MC 7+ , membrane-cytoplasmic linker plus seven C-terminal amino acids into the AAA+ ATPase module boundary; TM1, Transmembrane helix 1; TM2, Transmembrane helix 2. " width="100%" height="100%">
Journal: The Journal of Biological Chemistry
Article Title: The membrane-cytoplasmic linker defines activity of FtsH proteases in Pseudomonas aeruginosa clone C
doi: 10.1016/j.jbc.2023.105622
Figure Lengend Snippet: PaFtsH1 and PaFtsH2 have similar in vitro peptide-binding profiles and differentially affect in vivo colony size. A , peptide array of the RpoH sequence from Pseudomonas aeruginosa (PaRpoH; P42378|RpoH_PSEAE; 1–284, sliding window of 12 amino acids with a step size of two amino acids towards the C-terminus with each spot) probed with 1 μM 35 S-EcFtsH ( top ), 35 S-PaFtsH1 ( middle ), or 35 S-PaFtsH2 ( bottom ) in the presence of 1.25 mM ATPγS. Peptide sequences corresponding to each spot are listed in
Article Snippet: For protein induction, EcFtsH-Myc-6xHis, PaFtsH1-6xHis, and PaFtsH2-6xHis variants were expressed from a pET-28a(+) plasmid in
Techniques: In Vitro, Binding Assay, In Vivo, Peptide Microarray, Sequencing, Expressing, Plasmid Preparation, Derivative Assay, Clone Assay, Serial Dilution, Construct, Membrane

Journal: Scientific Reports
Article Title: Virulence and transcriptome profile of multidrug-resistant Escherichia coli from chicken
doi: 10.1038/s41598-017-07798-1
Figure Lengend Snippet: MICs (µg/mL) of five MDR E. coli isolates.
Article Snippet: In order to identify the molecular mechanism of virulent regulators in MDR E. coli , further transcriptome analysis of the most significant strain, E. coli 381, was performed and compared to the transcriptome profile of
Techniques:

Journal: Scientific Reports
Article Title: Virulence and transcriptome profile of multidrug-resistant Escherichia coli from chicken
doi: 10.1038/s41598-017-07798-1
Figure Lengend Snippet: Selected clinical isolates of poultry E. coli producing resistant genes.
Article Snippet: In order to identify the molecular mechanism of virulent regulators in MDR E. coli , further transcriptome analysis of the most significant strain, E. coli 381, was performed and compared to the transcriptome profile of
Techniques:

Journal: Scientific Reports
Article Title: Virulence and transcriptome profile of multidrug-resistant Escherichia coli from chicken
doi: 10.1038/s41598-017-07798-1
Figure Lengend Snippet: Biofilm formation of five isolates of E. coli and reference strain at different time intervals. The results are shown in the form of mean biofilm formation index (BFI) of three independent repeats and compared to E. coli ATCC 25922. Statistical significance (P ≤ 0.05) was calculated using two-tailed t-test. *P- values of ≤ 0.05 was considered significant higher than control. # P values of ≤0.05 was considered significant lower than control. **P- values of ≤0.05 was considered significant higher than control and *.
Article Snippet: In order to identify the molecular mechanism of virulent regulators in MDR E. coli , further transcriptome analysis of the most significant strain, E. coli 381, was performed and compared to the transcriptome profile of
Techniques: Two Tailed Test, Control

Journal: Scientific Reports
Article Title: Virulence and transcriptome profile of multidrug-resistant Escherichia coli from chicken
doi: 10.1038/s41598-017-07798-1
Figure Lengend Snippet: In vitro virulence assay of five E. coli isolates along with ATCC 25922 in Caco-2 cells. ( a ) Number of adherent and internalized bacteria. The results are presented as log10 of the mean ± standard deviation (SD) CFU/mL of three independent repeats and compared to E. coli ATCC 25922. ( b ) Intracellular survival rate of E. coli isolates at different time intervals. The results are presented as log10 CFU/mL mean ± SD of survival rate. Asterisk (*) represents statistical significance (P ≤ 0.05) using two-tailed t-test.
Article Snippet: In order to identify the molecular mechanism of virulent regulators in MDR E. coli , further transcriptome analysis of the most significant strain, E. coli 381, was performed and compared to the transcriptome profile of
Techniques: In Vitro, Bacteria, Standard Deviation, Two Tailed Test

Journal: Scientific Reports
Article Title: Virulence and transcriptome profile of multidrug-resistant Escherichia coli from chicken
doi: 10.1038/s41598-017-07798-1
Figure Lengend Snippet: In vitro virulence assay of five E. coli isolates along with ATCC 25922 in macrophage RAW 264.7 cells. ( a ) Number of adherent and internalized bacteria. The results are presented as log10 of the mean ± standard deviation (SD) CFU/mL of three independent repeats and compared to E. coli ATCC 25922. ( b ) Intracellular survival rate of E. coli isolates at different time intervals. The results are presented as log10 CFU/mL mean ± SD of survival rate. Asterisk (*) represents statistical significance (P ≤ 0.05) using two-tailed t-test.
Article Snippet: In order to identify the molecular mechanism of virulent regulators in MDR E. coli , further transcriptome analysis of the most significant strain, E. coli 381, was performed and compared to the transcriptome profile of
Techniques: In Vitro, Bacteria, Standard Deviation, Two Tailed Test

Journal: Scientific Reports
Article Title: Virulence and transcriptome profile of multidrug-resistant Escherichia coli from chicken
doi: 10.1038/s41598-017-07798-1
Figure Lengend Snippet: Determination of LD50 of the three multidrug resistant E. coli isolates in two-day old chickens.
Article Snippet: In order to identify the molecular mechanism of virulent regulators in MDR E. coli , further transcriptome analysis of the most significant strain, E. coli 381, was performed and compared to the transcriptome profile of
Techniques:

Journal: Scientific Reports
Article Title: Virulence and transcriptome profile of multidrug-resistant Escherichia coli from chicken
doi: 10.1038/s41598-017-07798-1
Figure Lengend Snippet: Important up-regulated and dwown-regulated genes in E. coli 381.
Article Snippet: In order to identify the molecular mechanism of virulent regulators in MDR E. coli , further transcriptome analysis of the most significant strain, E. coli 381, was performed and compared to the transcriptome profile of
Techniques: Conjugation Assay, Membrane, Binding Assay, Activation Assay, Chemotaxis Assay

Journal: Scientific Reports
Article Title: Virulence and transcriptome profile of multidrug-resistant Escherichia coli from chicken
doi: 10.1038/s41598-017-07798-1
Figure Lengend Snippet: Proposed resistance ( a ) and virulence ( b ) regulating genes and associated factors in isolate E. coli 381. Up-regulated genes and factors are shown with red-colored upward arrows while down-regulated genes and factors are shown with green-colored downward arrows.
Article Snippet: In order to identify the molecular mechanism of virulent regulators in MDR E. coli , further transcriptome analysis of the most significant strain, E. coli 381, was performed and compared to the transcriptome profile of
Techniques:

Journal: Cell stem cell
Article Title: Glucose modulates IRF6 transcription factor dimerization to enable epidermal differentiation.
doi: 10.1016/j.stem.2025.02.017
Figure Lengend Snippet: Figure 1. Intracellular glucose accumulates in keratinocyte differentiation (A) Heatmap representing average fold change of 614 metabolites identified in metabolomics, in early (day 3) and late (day 6) differentiation compared with the undifferentiated state (n = 5 biological replicates). (B) Accumulation of 2-NBDG (green), a fluorescent glucose analog, in differentiated keratinocytes with Hoechst nuclear stain (blue). Progenitor (PG) represents undifferentiated cells versus days 3 and 6 of calcium-induced differentiation in vitro (representative image of 3 independent samples, scale bar, 10 mm). (C and D) Quantitation of glucose-14C (C6) (C) or glucose-14C (C1) (D) accumulation during differentiation in culture, after 7 day incubation, normalized to mg of protein per sample (n = 3 biological replicates, 3 technical replicates per experiment). (E) Quantitation of glucose concentrations in regenerated epidermal organoid tissue, and whole epidermis undergoing stratification through day 6 analyzed (n = 3 biological replicates). (F) Epidermal organoids expressing vector control or the Red Glifon 3000 glucose sensor, and the dsRed signal (red) indicates detection of free glucose, with nuclear DAPI (blue) and phalloidin (green) stains (representative image of 3 biological replicates, scale bar, 50 mm). (G) Quantitation of dsRed intensity in the basal and spinous layers of the tissue normalized to phalloidin (n = 8 fields of view across 2 biological replicates). (H) Epidermis from transgenic mice expressing Red Glifon 3000; detection of free glucose (red), nuclear DAPI stain (blue), and immunostaining of differentiation marker keratin 10 (green); and gray bar indicates region of interest (ROI) used for quantitation in (I) (representative image shown, n = 3 biological replicates, scale bar, 50 mm). (I) Average fluorescence intensity with standard error (shaded region) in the ROI of Red Glifon, DAPI, and K10 across the epidermis from the basal layer up to but excluding the stratum corneum (n = 6). Data are represented as mean ± SEM, two-tailed unpaired Student’s t test was used for statistical analysis, *p < 0.05, ****p < 0.001. See also Figure S1 and Table S1.
Article Snippet: The iGlucoSnFR sensor (a gift from J. Keller and L. Looger, Addgene plasmid # 164514) and
Techniques: Staining, In Vitro, Quantitation Assay, Incubation, Expressing, Plasmid Preparation, Control, Transgenic Assay, Immunostaining, Marker, Two Tailed Test

Journal: Nature Communications
Article Title: Efficacy of modified-vaccinia Ankara vaccine as pre- and post-exposure prophylaxis against monkeypox sexual transmission in non-human primate model
doi: 10.1038/s41467-025-62681-2
Figure Lengend Snippet: Four cynomolgus macaques were challenged with MPXV IIb virus on day 0 with 1 × 10 7 pfu of MPXV by the rectal route. Two animals were euthanized on day 11 and two were followed for 83 days for clinical and immunological monitoring ( a ). Five clinical signs were monitored for 34 days ( n = 2): the presence of rectal exudate, loss of appetite, presence of lesions, lymphadenomegaly, and weight loss >10% were recorded for each animal. b Viral load assessment in rectal and seminal fluids ( n = 4 for rectal fluid and n = 2 for seminal plasma). MPXV DNA was quantified by qPCR and is expressed as viral DNA copies/mL. The lower limit of detection (LLOD) was 90,000 copies/mL, and the lower limit of quantification (LLOQ) was 100,000 copies/mL. c The rectal and seminal fluid samples containing viral DNA were tested for virus infectivity in Vero cells. Viral infectivity is quantified in pfu/mL. The LLOD was established at 10 PFU/mL, indicated by the dotted line. All samples below this LLOD were considered negative and are represented by a value of 1. d Orthopoxvirus-binding IgG concentrations are reported in arbitrary units (arb. unit./mL). Serum (left) and seminal plasma (right) from two NHPs ( n = 2) were evaluated for the total anti-MPXV protein IgG concentration before challenge and at various time points up to days 62 and 83 post-challenge for serum and seminal plasma, respectively. Each symbol represents one animal, and the different shapes represent different antigens. The dashes represent the mean. Ag: antigen. Source data are provided as a Source Data file.
Article Snippet:
Techniques: Virus, Clinical Proteomics, Infection, Binding Assay, Concentration Assay

Journal: Nature Communications
Article Title: Efficacy of modified-vaccinia Ankara vaccine as pre- and post-exposure prophylaxis against monkeypox sexual transmission in non-human primate model
doi: 10.1038/s41467-025-62681-2
Figure Lengend Snippet: Representative Hematoxylin-Eosin staining of the rectal mucosa on day 11 post-challenge from a cynomolgus macaque infected with MPXV. The analysis was performed on 2 animals. a Ulcerated rectal mucosa at the margin of the anus showing ulcerative, necrotizing rectitis (*) characterized by b hyperplastic mucosa (arrowhead) and c fibrin and extravasated erythrocytes (hemorrhage) (arrow head). d rectal submucosa, with numerous inflammatory cells (arrow head), mostly e neutrophils (calprotectin staining), f T cells (CD3 staining), g follicular-like B-cell aggregates (CD20 staining), and h minimal diffuse macrophage infiltration (CD68 staining). i RNA scope using a specific MPXV probe in the submucosal ulcerated tissue and adjacent epithelium (purple staining localized mostly in the ulcerated area). j RNAscope with the negative control probe DapB. Scale-bars: a = 500 µm, b – j = 100 µm.
Article Snippet:
Techniques: Staining, Infection, RNAscope, Negative Control

Journal: Nature Communications
Article Title: Efficacy of modified-vaccinia Ankara vaccine as pre- and post-exposure prophylaxis against monkeypox sexual transmission in non-human primate model
doi: 10.1038/s41467-025-62681-2
Figure Lengend Snippet: a Study design on NHP immunization and challenge periods in weeks, and a description of the four groups of cynomolgus macaques. One group was vaccinated before the challenge with two doses of MVA administered 4 weeks apart (MVA-PrEP, n = 4; gray). Another group was vaccinated with a single dose four days after the MPXV IIb challenge (MVA-PEP, n = 4; black). In addition, we challenged one group of convalescent NHPs previously exposed via the rectal or ID route alone or in combination 37–46 weeks before the challenge and described previously (Conv., n = 6; orange) and one control group (CTRL, n = 4; red). The four groups were challenged with 1 × 10 7 pfu of MPXV by the rectal route as indicated in blue. MVA-PrEP animals were monitored during the MVA immunization phase, with blood samplings for immunogenicity analysis. Following the MPXV challenge, blood, rectal fluid, seminal plasma, and skin swab samples were collected. The blue vertical dotted line represents the challenge. The image was created in BioRender. Herate, C. (2025) https://BioRender.com/lkzvios . b Rectal fluids collected during the challenge phase were evaluated for viral DNA content by qPCR. Viral load is expressed as viral DNA copies/mL. Each graph represents the viral kinetics of one group during 27 days. The LLOD was 36,000 copies/mL and the LLOQ 100,000 copies/mL. c Rectal fluid containing viral DNA was tested for virus infectivity in Vero cells and measured in pfu/mL. The LLOD was established at 10 PFU/mL, indicated by the dotted line. All samples below the LLOD were considered negative and are represented by a value of 1. When qPCR was negative, samples were considered negative and are represented by a value of 1. d Area under the curve (AUC) of viral DNA in rectal fluids during the 27 days of follow-up and statistical analysis. (CTRL, n = 4; red), (MVA-PEP, n = 4; black) (MVA-PrEP, n = 4; gray) (Conv., n = 6; orange). **: p = 0.0079. Each circle represents one animal. e Viral load measured by qPCR in skin swabs (left) and seminal plasma (right) sampled at day 11 and statistical analysis. The mean value is represented by the horizontal bar. (CTRL, n = 4 for skin and seminal fluid; red), (MVA-PEP, n = 3 for skin and n = 4 for seminal fluid; black) (MVA-PrEP, n = 3 for skin and n = 4 for seminal fluid; gray) (Conv., n = 5 for skin and n = 6 for seminal fluid; orange). *: p = 0.0333. Each circle represents one animal. Statistical analyses were performed using the Kruskal-Wallis test followed by a two-tailed non-parametric Mann-Whitney test (* p < 0.05, ** p < 0.01). Source data are provided as a Source Data file.
Article Snippet:
Techniques: Control, Immunopeptidomics, Clinical Proteomics, Virus, Infection, Two Tailed Test, MANN-WHITNEY

Journal: Nature Communications
Article Title: Efficacy of modified-vaccinia Ankara vaccine as pre- and post-exposure prophylaxis against monkeypox sexual transmission in non-human primate model
doi: 10.1038/s41467-025-62681-2
Figure Lengend Snippet: a The concentrations of five MPXV (upper panel) and homologous VACV (lower panel) protein-binding IgG were quantified in the serum of NHPs that were either immunized with MVA and/or challenged with MPXV. The data are expressed in arbitrary units (arb. unit./mL). Total IgG concentrations in NHP serum were monitored during the immunization phase for the MVA-PrEP group and after the challenge phase. For the remaining three groups, IgG follow-up began two weeks prior to the challenge. b Seroneutralization titers were measured in the serum collected at week -1 and week 4 post-challenge using Vero cells and live MPXV or VACV. The mean PRNT50 with standard deviation were calculated using infectious MPXV IIb (left panel): *: p = 0.0159, MPXV Ia (middle panel): *: p = 0.0159 or VACV-107 (right panel): *: p = 0.0190 and **: p = 0.0095. (CTRL, n = 4; red), (MVA-PEP, n = 4; black), (MVA-PrEP, n = 4; gray), (Conv., n = 6 at W-1 and n = 5 at W4; orange). c , d Cellular immunity conferred by MVA vaccination and/or MPXV challenge. (CTRL, n = 4; red), (MVA-PEP, n = 4; black) (MVA-PrEP, n = 4; gray) (Conv., n = 6 and n = 5 from W14; orange). Intracellular staining of PBMCs, collected during the MVA immunization phase and after MPXV challenge, after an overnight MPXV or MVA stimulation in vitro. Panels represent the mean percentage with standard deviation of CD4 + CD154 + cells expressing IFN-γ ( c ) and IL17A ( d ) upon MPXV 2b (left) or MVA (right) stimulation. For ( b – d ), each circle represents one animal. When represented, statistical analyses were performed using the Kruskal-Wallis test followed by a two-tailed non-parametric Mann-Whitney test (* p < 0.05, ** p < 0.01). Ag: antigen; Stim. Stimulation. Source data are provided as a Source Data file.
Article Snippet:
Techniques: Protein Binding, Standard Deviation, Staining, In Vitro, Expressing, Two Tailed Test, MANN-WHITNEY

Journal: Nature Communications
Article Title: Efficacy of modified-vaccinia Ankara vaccine as pre- and post-exposure prophylaxis against monkeypox sexual transmission in non-human primate model
doi: 10.1038/s41467-025-62681-2
Figure Lengend Snippet: CD4+ and CD8+ cells producing TNF-α, IL-2, and IFN-γ are represented for each animal (one pie chart per animal) at W12 post-immunization (W−1 post-challenge) and W15 post-immunization (W2 post-challenge). The polyfunctionality of CD4 + CD154 + (left) and CD8 + CD137 + (right) cells upon MPXV IIb stimulation is shown. Non-responder cells are not represented. Cells secreting only one of the three cytokines are in blue, two of them in green, and three in orange. Source data are provided as a Source Data file.
Article Snippet:
Techniques:

Journal: Nature Communications
Article Title: Efficacy of modified-vaccinia Ankara vaccine as pre- and post-exposure prophylaxis against monkeypox sexual transmission in non-human primate model
doi: 10.1038/s41467-025-62681-2
Figure Lengend Snippet: Correlation matrix between virological parameters measured after the MPXV challenge and humoral and cellular parameters measured 1 week before the challenge. Spearman's r values from −0.7 to −1 and from 0.7 to 1 are shown. Fl.: fluid.
Article Snippet:
Techniques:

Journal: Gut Microbes
Article Title: The Mucosally-Adherent Rectal Microbiota Contains Features Unique to Alcohol-Related Cirrhosis
doi: 10.1080/19490976.2021.1987781
Figure Lengend Snippet: Beta-diversity of EtOH vs non-EtOH cirrhosis with linear model of taxonomic changes. (a) Principal Coordinates Analysis via Bray–Curtis distances of rectal swabs by EtOH vs. non-EtOH etiology of cirrhosis ( p = .19; R 2 = 0.03). (b) Principal Coordinates Analysis via Bray–Curtis distances of stool by EtOH vs. non-EtOH etiology of cirrhosis ( p = .31; R 2 = 0.03). (c) Relative abundances of E. coli and Enterobacteriaceae in rectal swabs of EtOH vs. non-EtOH cirrhosis (FDR < 0.05). Gray dotted line indicates separation of high and low E. coli /Enterobacteriaceae groups
Article Snippet: To test this hypothesis, we incubated
Techniques:

Journal: Gut Microbes
Article Title: The Mucosally-Adherent Rectal Microbiota Contains Features Unique to Alcohol-Related Cirrhosis
doi: 10.1080/19490976.2021.1987781
Figure Lengend Snippet: E. coli growth in vitro under physiological concentrations of ethanol after alcohol consumption and its oxidative metabolites at pH 6. Growth curve in LB (aerobic, 37°C) with (a) ethanol, (b) acetaldehyde, (c) acetate, and (d) ethanol, acetaldehyde, and acetate alone and in combination. Data expressed as mean ± SEM. N = 3 replicates per condition
Article Snippet: To test this hypothesis, we incubated
Techniques: In Vitro

Journal: Nature Communications
Article Title: Cyclophilin J limits inflammation through the blockage of ubiquitin chain sensing
doi: 10.1038/s41467-018-06756-3
Figure Lengend Snippet: The C-terminal of CYPJ interacts with TAB2 and TAB3. a Effects of CYPJ on NF-κB reporter activation induced by individually transfected components of the NF-κB signal pathway ( N = 3). b , c Reciprocal co-IP assays determine the interaction between HA-CYPJ and Flag-TAB2 or Flag-TAB3 in 293T cells. d Endogenous co-IP assays evaluate the interaction between CYPJ and TAB2/TAB3 with or without TNF treatment for different time points in 293T cells. e Immunofluorescence microscopy observing the colocalization of pEGFP-CYPJ and Flag-TAB2 in HeLa cells. Scale bars, 20 μm. f Multiple sequence alignments of CYPJ proteins from different species. The three fragments of CYPJ are marked. g GST pull-down assays were performed using E.coli -expressed GST and GST-CYPJ fragments and lysates of 293T cells transfected with Flag-TAB2 or Flag-TAB3. h Co-IP assays detecting the interaction between Flag-TAB2/3 with HA-tagged FL or fragment-3 deleted CYPJ, # indicates a nonspecific band. Error bars indicate S.D.; n.s. no significance, * p < 0.05, ** p < 0.01 (two-tailed Student’s t -test)
Article Snippet: GST and GST fusion proteins were expressed and purified from
Techniques: Activation Assay, Transfection, Co-Immunoprecipitation Assay, Immunofluorescence, Microscopy, Sequencing, Two Tailed Test

Journal: Nature Communications
Article Title: Cyclophilin J limits inflammation through the blockage of ubiquitin chain sensing
doi: 10.1038/s41467-018-06756-3
Figure Lengend Snippet: CYPJ binds to the NZF domain of TAB2 and TAB3. a Schematic diagram of FL and truncation variants of TAB2. b HA-CYPJ and Flag-tagged TAB2 fragments were co-transfected into 293T cells for co-IP assays to detect their interaction. c GST pull-down assays were performed using E.coli -expressed GST and GST-TAB2 truncations and lysates of 293T cells transfected with HA-CYPJ. # indicates the bands of GST-fused full-length and truncated TAB2. d E.coli -expressed GST and GST-fused NZF domains of TAB2, TAB3, and NEMO were used in GST pull-down assays with lysates of 293T cells transfected with HA-CYPJ or HA-CYPA. e E.coli -expressed GST and GST-fused NZF domains of LUBAC members, as well as TAB2 and TAB3 were used in GST pull-down assays with E.coli -expressed His-CYPJ. f ITC assays were performed to detect the binding of in vitro synthesized NZF peptides derived from TAB2 or TAB3 to prokaryotically expressed His-CYPJ
Article Snippet: GST and GST fusion proteins were expressed and purified from
Techniques: Transfection, Co-Immunoprecipitation Assay, Binding Assay, In Vitro, Synthesized, Derivative Assay

Journal: bioRxiv
Article Title: Helicobacter small RNA regulates host adaptation and carcinogenesis
doi: 10.1101/2020.02.15.950279
Figure Lengend Snippet: a , Experimental strategies schematic. b , Circular genomic map of ATCC 43504 strain recovered from stomachs of gerbils and mice. c , Mutation rates and expression levels of candidate RNAs (mRNA or non-coding RNA). RNA expression levels of the ORFs or nearby genes of genome regions [total 15 genes , which mutated in more than 50% of the recovered strains from the gerbils, were assessed and plotted against the mutation rates. d , Schematic structures of genes around HPnc4160. e , Schematic diagram of genome DNA sequence around the HPnc4160 and poly-T sequence of recovered strains harboring mutations. Green indicates HPnc4160 transcribed sequence, red frame indicates poly-T mutated stretches in the upstream region of HPnc4160 of ATCC 43504 wild-type (WT) strain, and red-colored “T” indicated inserted nucleotides of each recovered strain compared with WT. f , The T-repeat length in the upstream region of HPnc4160 of clinical isolates. Isolated strains from cancer patients (Cancer) have a higher number of T-repeat in the upstream region of Hpnc4160 compared with the isolates from non-cancer patients (Non-Cancer). Data are presented as means with 95% confidence interval. P values represent the results of the two-tailed Mann-Whitney test. g , Expression levels of HPnc4160 in the recovered H. pylori strains from Mongolian gerbils (n=40). Relative expression levels of HPnc4160 were measured by real-time PCR and plotted against T-repeat length in the upstream region of HPnc4160. Data are presented as means with s.d. P values represent the results of two-tailed Dunn’s multiple comparison test. h , The relative expression levels of HPnc4160 in the H. pylori strains genetically modified with the T-repeat length. Data are presented as means with s.d. (n=3). P values are from Dunnett’s multiple comparison test (at two-sided).
Article Snippet: The
Techniques: Mutagenesis, Expressing, RNA Expression, Sequencing, Isolation, Two Tailed Test, MANN-WHITNEY, Real-time Polymerase Chain Reaction, Comparison, Genetically Modified

Journal: bioRxiv
Article Title: Helicobacter small RNA regulates host adaptation and carcinogenesis
doi: 10.1101/2020.02.15.950279
Figure Lengend Snippet: a, The list of mutation rates and expression levels of the candidate RNAs (mRNA or non-coding RNA) of strains recovered from stomachs of gerbils or mice 8 wks post-infection. The Locus tags highlighted in red indicated the candidates common in both of the strains originated from gerbils and mice. N/A, not applicable. b, The expression levels of candidate RNAs (mRNA or non-coding RNA) of isolates recovered from H. pylori ATCC43504-infected C57BL/6 mice stomachs. RNA expression levels of the genes or nearby genes of genome regions ( and ), which mutated in 100% of the recovered strains were assessed.
Article Snippet: The
Techniques: Mutagenesis, Expressing, Infection, RNA Expression

Journal: bioRxiv
Article Title: Helicobacter small RNA regulates host adaptation and carcinogenesis
doi: 10.1101/2020.02.15.950279
Figure Lengend Snippet: a, Schematic diagram of genome DNA sequence around the HPnc4160 and poly-T sequence of genome analyzed strains. Red-colored “T” indicated stretches of poly-T sequence. b, Schematic diagram of genome DNA sequence around the HPnc4160 and poly-T sequence of mice-recovered strains 8 wks after infection (n=10, Hp1 to Hp10). Green indicated HPnc4160 transcribed sequence, red frame indicated poly-T mutated stretches in the upstream region of HPnc4160, and red-colored “T” indicated inserted nucleotides of each recovered strain compared with wild-type. c-e, Time-dependent change in the length of the poly-T stretches upstream of the HPnc4160 coding region in gerbils or mice-recovered strains. c , Strains from Mongolian gerbils infected with ATCC 43504. d , Strains from C57BL/6 mice infected with ATCC 43504. e , Strains from C57BL/6 mice infected with PMSS1. Data are median with interquartile range. P values are from non-parametric Dunn’s multiple comparison test (at two-sided). ns: not significant. f , Expression levels of HPnc4160 in the recovered H. pylori strains from mice (n=10) and H. pylori wild-type (T-repeat 14). Relative expression levels of HPnc4160 were measured by real-time PCR and plotted against T-repeat length in the upstream region of HPnc4160. Data are presented as means with s.d. P values represent the results of two-tailed Dunn’s multiple comparison test.
Article Snippet: The
Techniques: Sequencing, Infection, Comparison, Expressing, Real-time Polymerase Chain Reaction, Two Tailed Test

Journal: bioRxiv
Article Title: Helicobacter small RNA regulates host adaptation and carcinogenesis
doi: 10.1101/2020.02.15.950279
Figure Lengend Snippet: a , Experimental strategies schematic. b , The raw data of the DNA sequence analysis of H. pylori genomes prepared from original culture (Original #1 - #3) and from passaged in vitro for 60 times (60-passaged #1 - #3).
Article Snippet: The
Techniques: Sequencing, In Vitro

Journal: bioRxiv
Article Title: Helicobacter small RNA regulates host adaptation and carcinogenesis
doi: 10.1101/2020.02.15.950279
Figure Lengend Snippet: a , Growth curves of H. pylori ATCC43504 mutants mutated in the number of T repeat in HPnc4160 upstream region. b-c, The relative expression levels of hpnc4170 ( b ) and HP0811 ( c ) in the H. pylori strains genetically modified with the T-repeat length. Data are presented as means with s.d. (n=3). P values are from Dunnett’s multiple comparison test (at two-sided). ns: not significant. d , The relative RNA expression levels of target candidates of HPnc4160 showed an inverse correlation with HPnc4160. The total RNA from the indicated H. pylori ATCC 43504 strains were extracted, reverse transcribed, and provided for qPCR to assess the indicated genes. The results represent the average of three separate experiments (each n=3). Data are presented as means ± s.d. (error bars). Spearman correlation coefficients (r) were used to evaluate the relationships among relative RNA expression of HPnc4160 and each target.
Article Snippet: The
Techniques: Expressing, Genetically Modified, Comparison, RNA Expression, Reverse Transcription
![a , The MA plot of ratios [Δ hpnc4160 / hpnc4170 / wild-type (WT) H. pylori ] versus their normalized average mRNA expression determined by RNA-sequencing (RNA-Seq). The red dots showed genes of P <0.001. b , Volcano plots of the proteins quantified by isobaric tags for relative and absolute quantification (iTRAQ) analysis comparing WT and Δ hpnc4160 / 4170 . Each point represents the difference in expression (fold change) between the two groups plotted against the level of statistical significance. The red dots showed proteins of P <0.01. c , The Venn diagram represents the number of factors whose expression exhibited significant differences between Δ hpnc4160 / hpnc4170 and WT. d , The relative RNA expression levels of target candidates of HPnc4160 showed an inverse correlation with HPnc4160. The results represent the average of three separate experiments (each n=3). Data are presented as means ± s.d. (error bars). P values are from Tukey’s multiple comparison test (at two-sided). ns: not significant.](https://bio-rxiv-images-cdn.bioz.com/dois_ending_with_79/10__1101_slash_2020__02__15__950279/10__1101_slash_2020__02__15__950279___F2.large.jpg)
Journal: bioRxiv
Article Title: Helicobacter small RNA regulates host adaptation and carcinogenesis
doi: 10.1101/2020.02.15.950279
Figure Lengend Snippet: a , The MA plot of ratios [Δ hpnc4160 / hpnc4170 / wild-type (WT) H. pylori ] versus their normalized average mRNA expression determined by RNA-sequencing (RNA-Seq). The red dots showed genes of P <0.001. b , Volcano plots of the proteins quantified by isobaric tags for relative and absolute quantification (iTRAQ) analysis comparing WT and Δ hpnc4160 / 4170 . Each point represents the difference in expression (fold change) between the two groups plotted against the level of statistical significance. The red dots showed proteins of P <0.01. c , The Venn diagram represents the number of factors whose expression exhibited significant differences between Δ hpnc4160 / hpnc4170 and WT. d , The relative RNA expression levels of target candidates of HPnc4160 showed an inverse correlation with HPnc4160. The results represent the average of three separate experiments (each n=3). Data are presented as means ± s.d. (error bars). P values are from Tukey’s multiple comparison test (at two-sided). ns: not significant.
Article Snippet: The
Techniques: Expressing, RNA Sequencing, Quantitative Proteomics, Multiplex sample analysis, RNA Expression, Comparison

Journal: bioRxiv
Article Title: Helicobacter small RNA regulates host adaptation and carcinogenesis
doi: 10.1101/2020.02.15.950279
Figure Lengend Snippet: a, Growth curves of H. pylori ATCC43504 mutants. b, Predicted the secondary structure of HPnc4160 RNA by CentroidFold. The bases in the predicted structure are colored according to base-pairing probabilities. Circles in pink and light green color indicated loop structures having probabilities of binding to target RNA sequences. c, Schematic diagram of predicted HPnc4160 binding sites in the corresponding 5’UTR sequence of target genes. Upper sequences indicate target mRNA sequences with base numbers, whereas lower sequences indicate HPnc4160 sequence. Colored sequences are corresponding to the loop structures indicated in ( b ). d , Binding prediction of HPnc4160 and 5’ UTR of cagA mRNA. d , Schematic diagram of predicted HPnc4160 binding sites in the cagA CM/CRPIA motif of cagA CDS. e-f, Schematic diagram of predicted HPnc4160 binding sites in the corresponding CDS sequence of cagA TYPE 1 ( e ), TYPE 2 ( f ), and cagA nonbinding form (NB- cagA ) of TYPE 2 ( g ) and TYPE 1 ( h ). Upper sequences indicate target cagA mRNA sequences, whereas lower sequences indicate HPnc4160 sequence with base numbers. Colored sequences are corresponding to the loop structures indicated in ( b ). Mutated nucleotides in cagA mRNA sequence are shown in red. i , Purified RNase III was separated by SDS-PAGE and stained with CBB.
Article Snippet: The
Techniques: Binding Assay, Sequencing, Purification, SDS Page, Staining

Journal: bioRxiv
Article Title: Helicobacter small RNA regulates host adaptation and carcinogenesis
doi: 10.1101/2020.02.15.950279
Figure Lengend Snippet: a, Growth curves of H. pylori ATCC43504 cagA -NB mutant compared with wild-type. b , Phosphorylated CagA protein levels in cell lysates of AGS cells infected with H. pylori ATCC43504. The whole-cell lysates of AGS cells infected with H. pylori strains for 6 hours were subjected to western blot against anti-CagA, anti-pY CagA, anti-UreA, and anti-Actin antibodies. The band intensities were measured and calculated by ImageJ software. c , IL-8 production from AGS cells infected with H. pylori ATCC43504. The supernatants from AGS cells infected with H. pylori strains shown in the figure for the indicated time were subjected to ELISA system for IL-8 production. The results represent the average of three separate experiments (each n=3). Data are presented as means ± s.d. (error bars).
Article Snippet: The
Techniques: Mutagenesis, Infection, Western Blot, Software, Enzyme-linked Immunosorbent Assay

Journal: bioRxiv
Article Title: Helicobacter small RNA regulates host adaptation and carcinogenesis
doi: 10.1101/2020.02.15.950279
Figure Lengend Snippet: a , HPnc4160, and cagA mRNA expression levels. The results represent the average of three separate experiments (each n=3). Data are presented as means ± s.d. (error bars). P values are from non-parametric Dunnett’s multiple comparison test (at two-sided). ns: not significant. Experiments were repeated three times with similar results. b , Protein expression levels of CagA in each mutant strain. UreA protein levels serve as bacterial loading controls. c and d , Scattering phenotypes of H. pylori -infected AGS gastric epithelial cells. c , DNA (blue), F-actin (red), and anti-phosphorylated CagA antibody (pY-CagA, green) were stained. Scale bar, 50 μm. d , Quantification of scattering activity of AGS cells induced by H. pylori infection. e and f , HPnc4160 deletion mutants efficiently colonized the stomach and contributed to increasing mRNA levels of inflammatory chemokine Cxcl2 in mice stomach. C57BL/6 mice were inoculated with H. pylori . At three days after infection, animals were sacrificed, and a quantitative culture assay ( e ) and a quantitative RT-PCR ( f ) were performed on gastric specimens. Data are median with interquartile range. P values are from non-parametric Dunn’s multiple comparison test (at two-sided). ns: not significant. g, Clinical isolates from malignant patients downregulate HPnc4160 and upregulate expression levels of its target genes. Expression levels of indicated mRNAs in clinical isolates of non-cancer (Non-Cancer, n=39) and cancer (Cancer, n=17) patients were quantified and normalized with the levels of 23S rRNA. Data are presented as medians with interquartile range. P values are from the non-parametric Mann-Whitney test (at two-sided).
Article Snippet: The
Techniques: Expressing, Comparison, Mutagenesis, Infection, Staining, Activity Assay, Quantitative RT-PCR, MANN-WHITNEY

Journal: bioRxiv
Article Title: Helicobacter small RNA regulates host adaptation and carcinogenesis
doi: 10.1101/2020.02.15.950279
Figure Lengend Snippet: H. pylori infection in vivo leads elongation of T-stretch in the upstream region of HPnc4160 sRNA coding region, which results in decreased expression levels of sRNA HPnc4160. Gene silencing of HPnc4160 results in increased levels of target genes coding OMPs and CagA, and as a result, the levels of bacterial colonization and CagA translocation into the attached host cells were increased.
Article Snippet: The
Techniques: Infection, In Vivo, Expressing, Translocation Assay

Journal: eLife
Article Title: Toxoplasma bradyzoites exhibit physiological plasticity of calcium and energy stores controlling motility and egress
doi: 10.7554/eLife.73011
Figure Lengend Snippet: ( A ) Egress of tachyzoites and bradyzoites in response to A23187 (2 μM) for 15 min. Anti-GRA7, anti-SAG1, and anti-BAG1 antibodies followed by secondary antibodies to Alexa conjugated fluorochromes were used to detect the parasitophorous vacuole (PV) membrane, tachyzoites, and bradyzoites, respectively. Dolichos biflorus agglutinin (DBA) conjugated to FITC was used to stain the cyst wall. Arrow indicates released bradyzoites. Scale bar = 10 μm. ( B ) Quantitative analysis of egress in response to A23187 (2 μM) or zaprinast (500 μM) in extracellular buffer (EC) with Ca 2+ for 15 min. Each data point represents the percentage of egressed or released parasites from one PV or cyst (n = 20). Means ± SD of two independent experiments with 20 replicates. Two-tailed Mann–Whitney test, ***p<0.001. ( C ) Quantitative analysis of maximum distance egressed or released parasites moved away from the vacuole/cyst in response to A23187 (2 μM) or zaprinast (500 μM) in EC buffer with Ca 2+ for 15 min. Each data point represents distance traveled of one egressed tachyzoite or released bradyzoite from the original PV or cyst (n = 20). Means ± SD of two independent experiments with 20 replicates. Two-tailed Mann–Whitney test, ***p<0.001. ( D ) Quantitative analysis of speed (μm/s) of egressed or released parasites in response to A23187 (2 μM) or zaprinast (500 μM) in EC buffer with Ca 2+ for 15 min by time-lapse microscopy. Mean speed was determined by time-lapse recording during the first 1 min after egress or release. Each data point represents migration speed of a single egressed tachyzoites or released bradyzoites from original PV or cyst (n = 20). Means ± SD of two independent experiments with 20 replicates. Two-tailed unpaired Student’s t -test, ***p<0.001. Figure 1—source data 1. Percentage of egress or release from one vacuole or cyst (related to ). Figure 1—source data 2. Maximum egress distance of egressed or released parasites (related to ). Figure 1—source data 3. Speed of egressed or released parasites (related to ).
Article Snippet: Cysts were released from host cells by repeated passage through a 23G needle and collected by centrifugation at 150 × g for 10 min. To purify large-scale bradyzoites used for western blotting, pelleted cysts were resuspended in 1 ml PBS containing 10 µl
Techniques: Staining, Two Tailed Test, MANN-WHITNEY, Time-lapse Microscopy, Migration

Journal: eLife
Article Title: Toxoplasma bradyzoites exhibit physiological plasticity of calcium and energy stores controlling motility and egress
doi: 10.7554/eLife.73011
Figure Lengend Snippet: ( A ) Schematic of bradyzoites MIC2 secretion assay using ME49 BAG1-mCherry MIC2-GLuc bradyzoites, differentiated in vitro by cultivation at pH 8.2 for 7 days, based on fluorescence-activated cell sorting (FACS). ( B ) Immunofluorescence assay (IFA) analysis showing localization of MIC2-Gluc in bradyzoites induced for 7 days at pH 8.2. MIC2-Gluc was stained with anti-Myc antibody, bradyzoites were detected with anti-mCherry, followed by secondary antibodies conjugated with Alexa Fluor dyes, and the cyst wall was stained with DBA-FITC. Bar = 5 μm. ( C ) Western blots showing the expression of MIC2-Gluc and M2AP in tachyzoites and bradyzoites (induced for 7 days at pH 8.2, purified by magnetic beads and released from in vitro cysts by 0.25 mg/ml trypsin) of ME49 BAG1-mCherry MIC2-GLuc reporter. αMyc and αM2AP antibodies were used to probe the expression of MIC2-GLuc and M2AP, respectively. SAG1 and BAG1 serve as the stage-specific markers of tachyzoites and bradyzoites, respectively. Actin was used as a loading control. ( D ) ME49 BAG1-mCherry MIC2-Gluc tachyzoites or bradyzoites sorted by FACS and resuspended in extracellular (EC) buffer with Ca 2+ were stimulated by 0.1% DMSO, ionomycin (1 μM), or zaprinast (500 μM) for 10 min at 37°C. Release of MIC2-GLuc in excretory-secretory antigens (ESA) was determined using a Gaussia luciferase assay. RLU indicates relative light units. Means ± SEM of three independent experiments each with three replicates. Multiple Student’s t -tests, ***p<0.001. ( E ) Schematic illustration of the FNR-mCherry BAG1-EGFP dual fluorescence reporter and leakage of FNR-mCherry from the parasitophorous vacuole (PV) (top) or cyst matrix (bottom) following A23187-induced membrane permeabilization. ( F ) FNR-mCherry leakage was monitored by time-lapse imaging of FNR-mCherry after A23187 (2 μM) treatment. FNR-mCherry BAG1-EGFP tachyzoites cultured under normal condition for 24 hr or bradyzoites induced for 7 days at pH 8.2 were treated with A23187 (2 μM) or 0.25 mg/ml trypsin in EC buffer with Ca 2+ for 10 min at 37℃. Dash circle indicates the region of interest (ROI) for measurement of fluorescence intensity. Bar = 10 μm. ( G ) FNR-mCherry fluorescence (F) over the initial signal (F 0 ) vs. time from cells treated as in ( F ). Curves are made of data from five independent vacuoles or cysts and shown as means ± SD. Bradyzoites treated with the DMSO group were used to assess photobleaching of mCherry (gray line). DBA, Dolichos biflorus agglutinin. Figure 2—source data 1. Western blotting analysis of ME49 BAG1-mCherry MIC2-GLuc reporter parasites (related to ). Figure 2—source data 2. Determining MIC2-Gluc secretion by parasites using Gaussia luciferase assay (related to ). Figure 2—source data 3. Monitoring leakage of FNR-mCherry from the parasitophorous vacuole (PV) or cyst matrix following A23187-induced membrane permeabilization (related to ).
Article Snippet: Cysts were released from host cells by repeated passage through a 23G needle and collected by centrifugation at 150 × g for 10 min. To purify large-scale bradyzoites used for western blotting, pelleted cysts were resuspended in 1 ml PBS containing 10 µl
Techniques: In Vitro, Fluorescence, FACS, Immunofluorescence, Staining, Western Blot, Expressing, Purification, Magnetic Beads, Luciferase, Imaging, Cell Culture

Journal: eLife
Article Title: Toxoplasma bradyzoites exhibit physiological plasticity of calcium and energy stores controlling motility and egress
doi: 10.7554/eLife.73011
Figure Lengend Snippet: ( A ) Immunofluorescence assay (IFA) analysis showing co-localization of MIC2-Gluc with M2AP in bradyzoites induced for 7 days at pH 8.2. MIC2-Gluc was stained with anti-Myc antibody, M2AP was detected with anti-M2AP, followed by secondary antibodies conjugated with Alexa Fluor dyes, and the cyst wall was stained with biotinylated Dolichos biflorus agglutinin (DBA), followed by Alexa Fluor 647 conjugated streptavidin. Bar = 5 μm. ( B ) Bradyzoites expressing BAG1-mCherry were induced for 7 days at pH 8.2, mechanically liberated from cysts by 0.25 mg/ml trypsin for 5 min in intracellular buffer (IC buffer), and collected by fluorescence-activated cell sorting (FACS) after gating with parental ME49 ∆ hxgprt::Fluc parasites.
Article Snippet: Cysts were released from host cells by repeated passage through a 23G needle and collected by centrifugation at 150 × g for 10 min. To purify large-scale bradyzoites used for western blotting, pelleted cysts were resuspended in 1 ml PBS containing 10 µl
Techniques: Immunofluorescence, Staining, Expressing, Fluorescence, FACS

Journal: bioRxiv
Article Title: Sequencing of N 6 -methyl-deoxyadenosine at single-base resolution across the mammalian genome
doi: 10.1101/2023.01.16.524325
Figure Lengend Snippet: A) The 6mA-2-thio-dTTP base pair forms weaker hydrogen bonds compared to dA-dTTP and dA-2-thio-dTTP base pairs. B) Heatmap plot for the 6mA-to-T/C/G mutation ratios at 256 motifs (NN-6mA-NN) on the single-stranded model DNA after mutation-prone primer extension. C) The mutation patterns (6mA to T, C, and G respectively) at 256 motifs (NN-6mA-NN) on the single-stranded model DNA. D) FTO demethylates over 99% methylated base of NN-6mA-NN model ssDNA in vitro and shows weaker activity on dsDNA. Data are mean ± s.e.m.; analyzed by two-tailed unpaired t-tests. The number of independently repeated reactions are shown in each plot. **** P <0.0001. E) Schematic diagram of DR-6mA-seq. Genomic DNA is sequentially denatured, mixed with spike-in probes, treated with FTO (or not), ligated to biotin-tagged adaptors, and extended by Bst 2.0 DNA polymerase or Q5 high-fidelity DNA polymerase. The synthesized DNA strand is enriched and amplified for next-generation sequencing. 6mA-to-T/C/G mutations were counted for defining 6mA sites and calculating 6mA methylation stoichiometry, using spike-in calibration curves. F) Mutation profiles (including 6mA to T, C, and G) of mutation-prone primer extension on the 6mA spike-in probes harboring 0%, 25%, 50%, 75% and 100% 6mA. Primer extension on FTO-treated probes and primer extension by Q5 high-fidelity DNA polymerase serve as two control groups and do not respond to 6mA fractions. See also Table S1 .
Article Snippet:
Techniques: Mutagenesis, Methylation, In Vitro, Activity Assay, Two Tailed Test, Synthesized, Amplification, Next-Generation Sequencing, Control
Journal: bioRxiv
Article Title: Sequencing of N 6 -methyl-deoxyadenosine at single-base resolution across the mammalian genome
doi: 10.1101/2023.01.16.524325
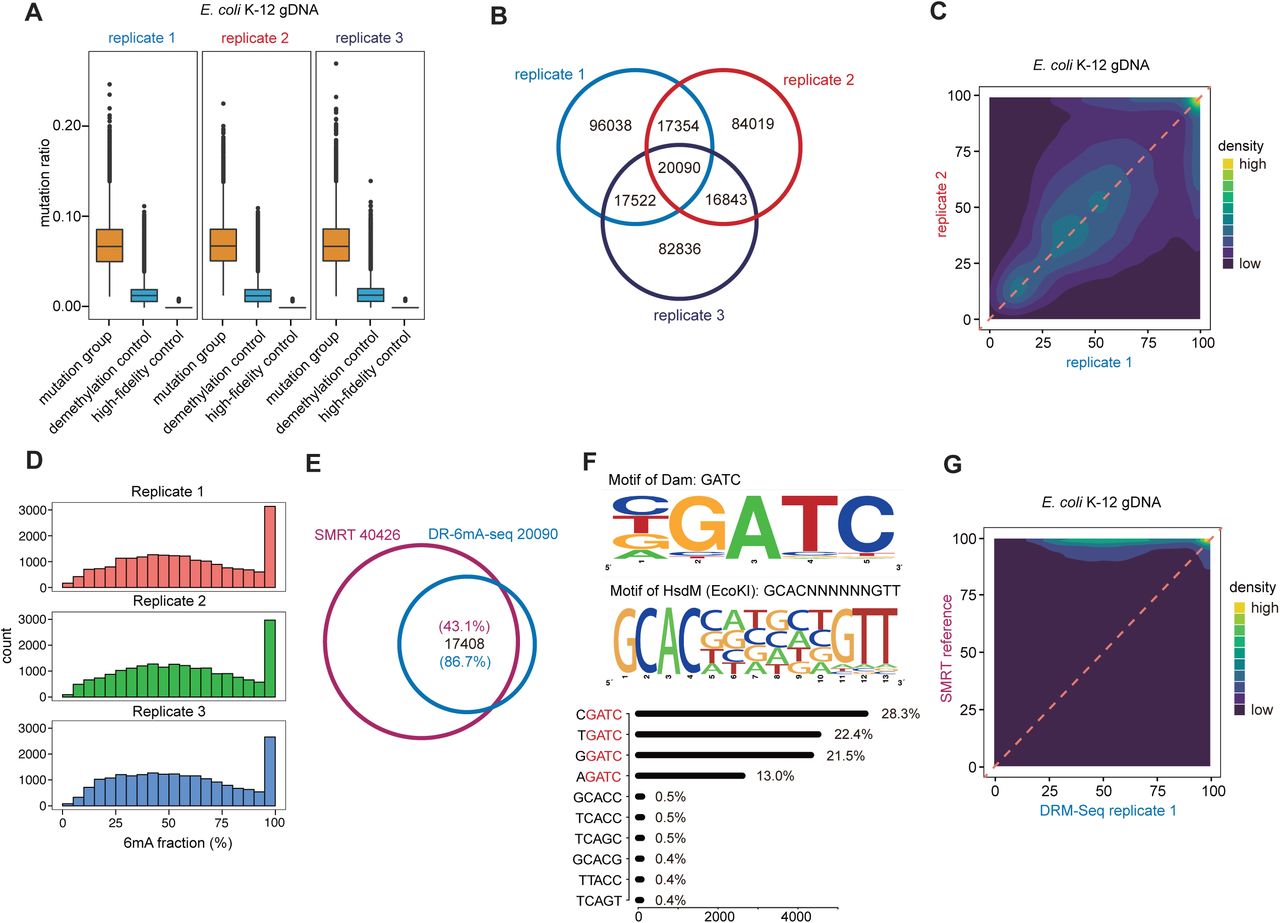
Figure Lengend Snippet: A) The box plot of mutation ratio distribution at 6mA sites detected in E. coli gDNA, revealed by DR-6mA-seq. The mutation ratios of three groups are shown, i.e., Bst 2.0 extended DNA from untreated DNA (mutation group), Bst 2.0 extended DNA from FTO-treated DNA (demethylation control), and Q5-extended DNA from untreated DNA (high-fidelity control). B) Venn diagram showing the overlapped 6mA sites among three biologically independent replicates, detected by DR-6mA-seq. C) Correlation analysis of methylation fractions in E. coli K-12 genome, indicating a high correlation between replicates of DR-6mA-seq. D) Histogram indicating the distribution of 6mA fractions at the modified sites in E. coli K-12 genome, revealed by DR-6mA-seq. E) Venn diagram showing the excellent overlap between DR-6mA-seq-detected 6mA sites and SMRT-detected 6mA sites in E. coli K-12 genome. F) Motif sequence logo and the list of consensus motifs containing 6mA sites in gDNA from E. coli K-12, uncovered by DR-6mA-seq. The frequency percentages of the top 5-base 6mA-containing motifs are shown. G) Correlation analysis of 6mA methylation fractions in E. coli K-12 genome, indicating a high correlation between 6mA sites detected by DR-6mA-seq and SMRT. See also Tables S2.
Article Snippet:
Techniques: Mutagenesis, Control, Methylation, Modification, Sequencing

Journal: bioRxiv
Article Title: Sequencing of N 6 -methyl-deoxyadenosine at single-base resolution across the mammalian genome
doi: 10.1101/2023.01.16.524325
Figure Lengend Snippet: A) The box plot of mutation ratio distribution at 6mA sites detected in HepG2 mtDNA, mouse brain mtDNA, and mouse liver mtDNA, revealed by DR-6mA-seq. The mutation ratios of three groups are shown, such as Bst 2.0 -extended DNA from untreated DNA (mutation group), Bst 2.0 -extended DNA from FTO-treated DNA (demethylation control), and Q5-extended DNA from untreated DNA (high-fidelity control). n = 2, biologically independent replicates. B) The histogram of mtDNA 6mA site number distribution normalized to 6mA fraction bins, in HepG2 cells, mouse brain, and mouse liver. n = 2, biologically independent replicates. C) Distribution of 6mA sites along the human HepG2 mitochondrial genome. The dots on the lighter orange background are mtDNA 6mA sites on the forward and reverse strand of mtDNA of HepG2 replicate 1, while the dots on the darker orange background are mtDNA 6mA sites on the forward and reverse strand of mtDNA of HepG2 replicate 2, both revealed by DR-6mA-seq. The six circled axes representing the methylation fractions at 0%, 20%, 40%, 60%, 80%, and 100%. The inner bright orange track represents overlapped mtDNA 6mA sites with 5-bp flanking windows. The inner purple track represents 6mA sites detected by ChIP-exo. D) Distribution of 6mA sites along the mouse mitochondrial genome. The dots on the purple and green background are mtDNA 6mA sites in two biological replicates of mouse brain and mouse liver, respectively, revealed by DR-6mA-seq, with six circled axes representing the methylation fractions at 0%, 20%, 40%, 60%, 80%, and 100%. The inner red track indicates the regions of uniquely mapped reads in mouse mitochondria genome. E) Chromatogram of a representative injection of 6mA (0.001 pmol) and dG standard compound (800 pmol), using LC-MS/MS. The 6mA quantification was calculated by the 6mA peak area, which displays a retention time of ∼5.8 min. F) Chromatogram of a representative injection of 6mA (0.005 pmol) and dG standard compound (300 pmol) using LC-MS/MS. The 6mA quantification was calculated by the 6mA peak area, which displays a retention time of ∼5.8 min. G) Chromatogram of a representative injection of gDNA-depleted HEK293T mtDNA (the first biological replicate), using LC-MS/MS. The optimized LC protocol can distinguish the 6mA peak from the dT+Na + peak which appears at ∼4.2 min. H) Chromatogram of a representative injection of gDNA-depleted HEK293T mtDNA (the second biological replicate), using LC-MS/MS. The optimized LC protocol can distinguish the 6mA peak from the dT+Na + peak which appears at ∼4.2 min. I) Chromatogram of a representative injection of gDNA-depleted HEK293T mtDNA (the third biological replicate), using LC-MS/MS. The optimized LC protocol can distinguish the 6mA peak from the dT+Na + peak which appears at ∼4.2 min.
Article Snippet:
Techniques: Mutagenesis, Control, Methylation, Injection, Liquid Chromatography with Mass Spectroscopy
Journal: bioRxiv
Article Title: Sequencing of N 6 -methyl-deoxyadenosine at single-base resolution across the mammalian genome
doi: 10.1101/2023.01.16.524325
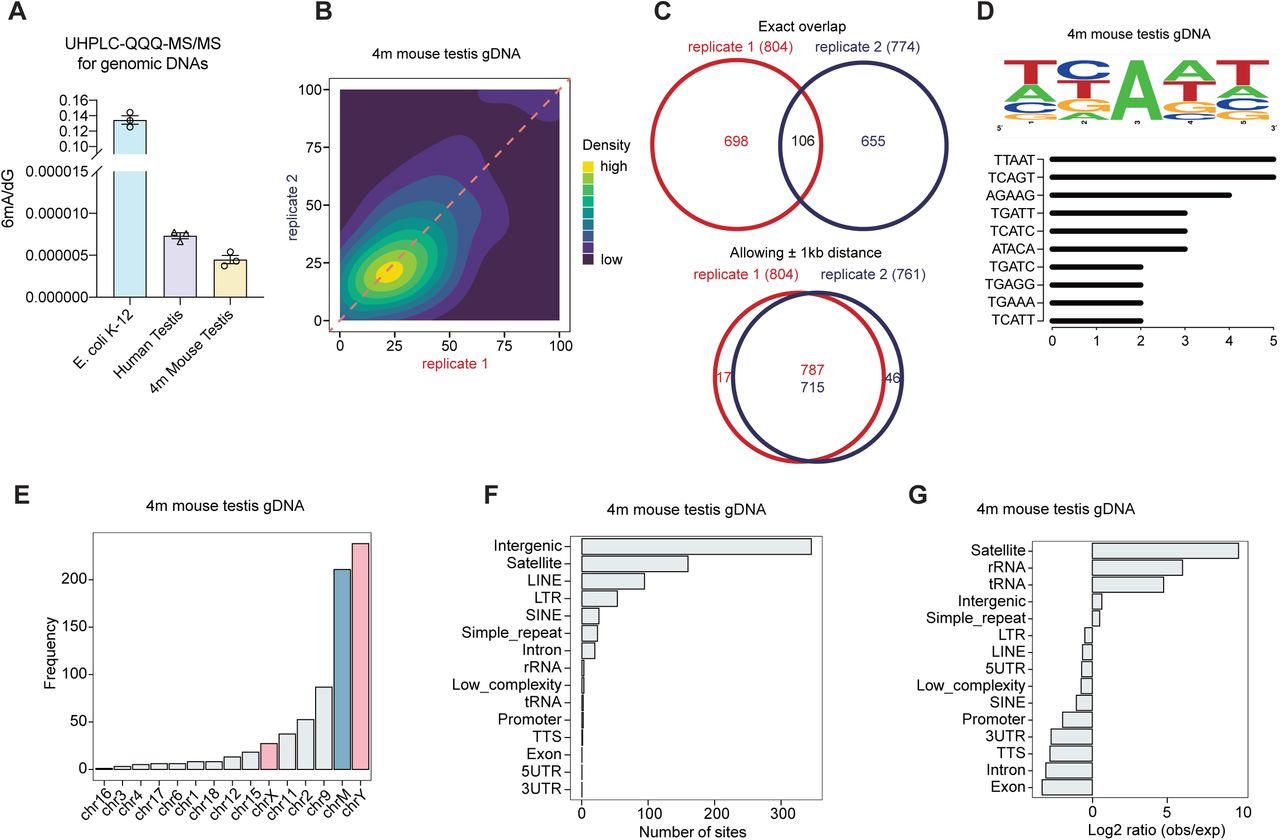
Figure Lengend Snippet: A) Quantification of 6mA level in gDNA from human and mouse intact testis by LC-MS/MS, with E. coli gDNA as a positive control. Data are mean ± s.e.m.. B) Correlation analysis of 6mA methylation fractions in antibody-enriched mouse testis gDNA, indicating a high correlation between replicates of DR-6mA-seq. C) Venn diagram showing the overlapped 6mA sites in mouse testis gDNA, allowing and not allowing ± 1 kb window, detected by DR-6mA-seq between two biologically independent replicates. D) Motif sequence logo and the list of the consensus motifs containing 6mA sites in gDNA from mouse testis, uncovered by DR-6mA-seq. E) Chromosome-wide distributions of gDNA 6mA sites in mouse testis, revealed by DR-6mA-seq. F) Genomic distributions of gDNA 6mA sites in mouse testis, revealed by DR-6mA-seq. G) Genomic enrichment of gDNA 6mA sites in mouse testis, revealed by DR-6mA-seq. See also , , and Tables S6.
Article Snippet:
Techniques: Liquid Chromatography with Mass Spectroscopy, Positive Control, Methylation, Sequencing
Journal: bioRxiv
Article Title: Sequencing of N 6 -methyl-deoxyadenosine at single-base resolution across the mammalian genome
doi: 10.1101/2023.01.16.524325
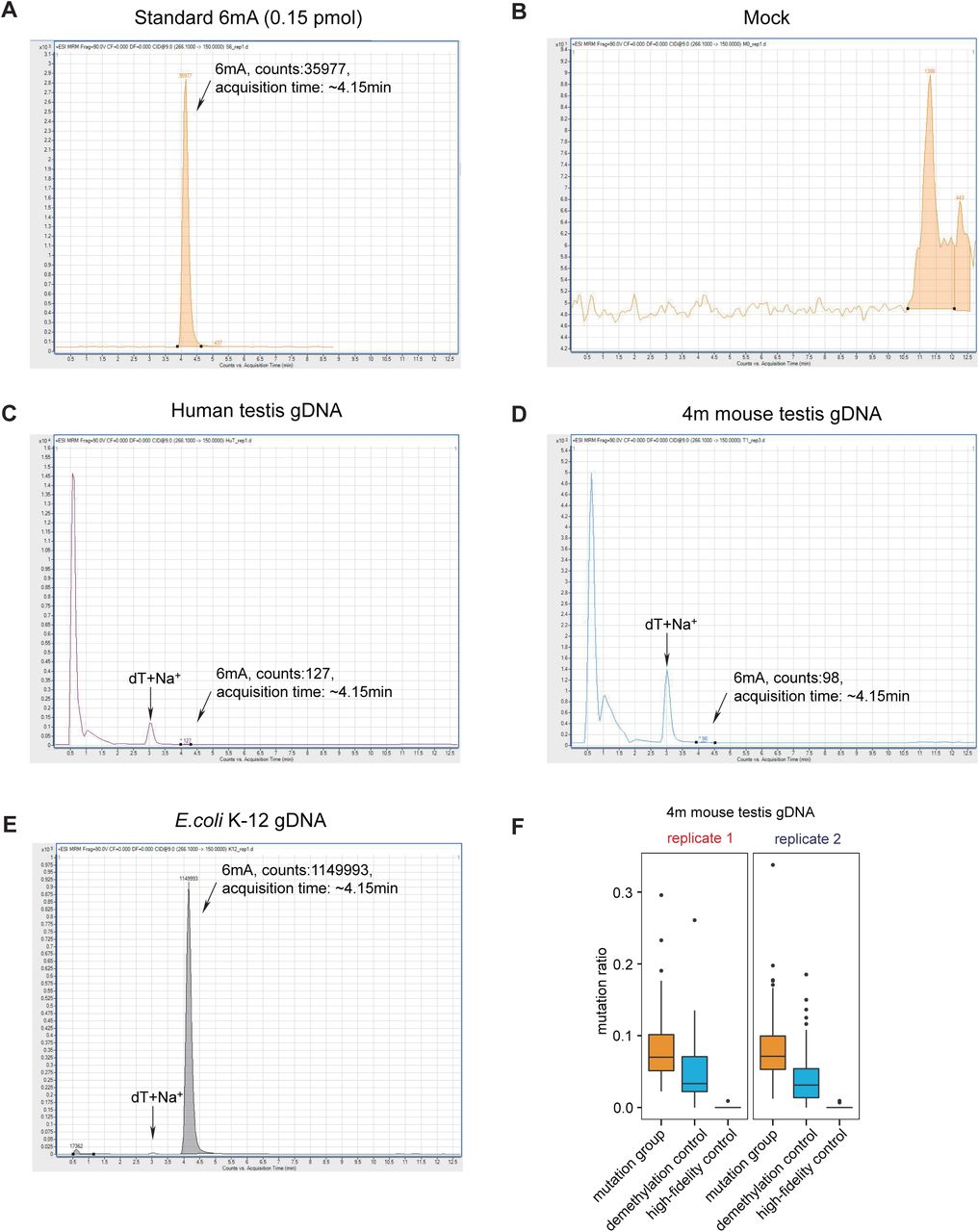
Figure Lengend Snippet: A) Chromatogram of a representative injection of 6mA standard compound, using LC-MS/MS. B) Chromatogram of a representative injection of the mock (nucleoside digestion mixture without adding any DNA) using LC-MS/MS. C) Chromatogram of a representative injection of digested human testis gDNA, using LC-MS/MS. The 6mA quantification was calculated by the 6mA peak area, which displays a retention time of ∼4.15 min. The optimized LC protocol distinguished the 6mA peak from the dT+Na + peak which appears at ∼3.00 min. D) Chromatogram of a representative injection of digested mouse testis gDNA, using LC-MS/MS. E) Chromatogram of a representative injection of E. coli K-12 gDNA, using LC-MS/MS. F) The box plot of mutation ratio distribution at 6mA sites detected in mouse testis gDNA, revealed by DR-6mA-seq. The mutation ratios of three groups are shown, such as Bst 2.0 -extended DNA from untreated DNA (mutation group), Bst 2.0 -extended DNA from FTO-treated DNA (demethylation control), and Q5-extended DNA from untreated DNA (high-fidelity control).
Article Snippet:
Techniques: Injection, Liquid Chromatography with Mass Spectroscopy, Mutagenesis, Control
Journal: bioRxiv
Article Title: Sequencing of N 6 -methyl-deoxyadenosine at single-base resolution across the mammalian genome
doi: 10.1101/2023.01.16.524325
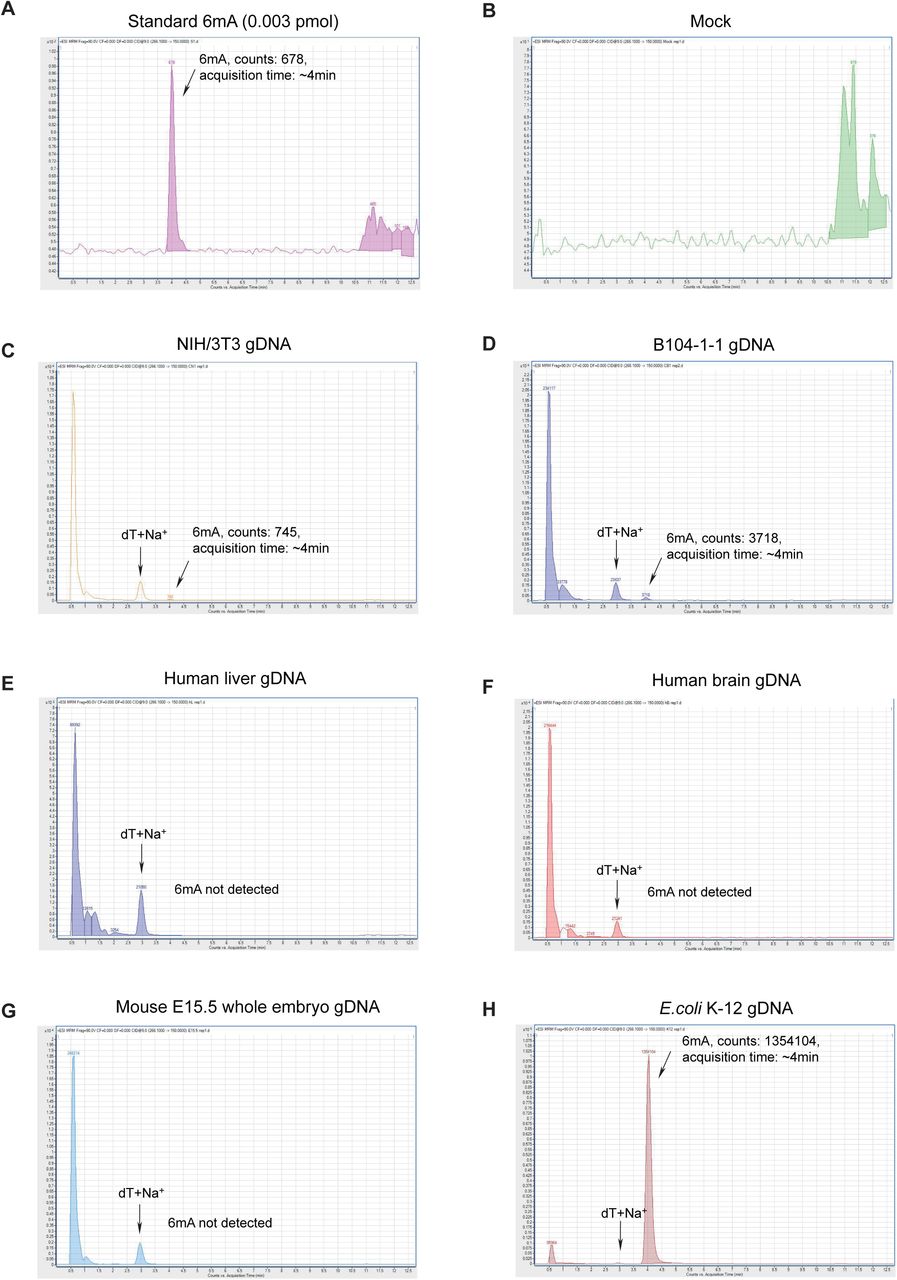
Figure Lengend Snippet: A) Chromatogram of a representative injection of 6mA standard compound, using LC-MS/MS. B) Chromatogram of a representative injection of the mock (nucleoside digestion mixture without adding any DNA) using LC-MS/MS. C) Chromatogram of a representative injection of digested gnotobiotic NIH/3T3 (ATCC) gDNA, using LC-MS/MS. The 6mA quantification was calculated by the 6mA peak area, which displays a retention time of ∼4.15 min. The optimized LC protocol distinguished the 6mA peak from the dT+Na + peak which appears at ∼3.00 min. D) Chromatogram of a representative injection of digested gnotobiotic B104-1-1 (ATCC) gDNA, using LC-MS/MS. E) Chromatogram of a representative injection of digested human liver gDNA, using LC-MS/MS. F) Chromatogram of a representative injection of digested human brain gDNA, using LC-MS/MS. G) Chromatogram of a representative injection of digested mouse E15.5 whole embryo gDNA, using LC-MS/MS. H) Chromatogram of a representative injection of E. coli K-12 gDNA, using LC-MS/MS.
Article Snippet:
Techniques: Injection, Liquid Chromatography with Mass Spectroscopy
Journal: bioRxiv
Article Title: Sequencing of N 6 -methyl-deoxyadenosine at single-base resolution across the mammalian genome
doi: 10.1101/2023.01.16.524325
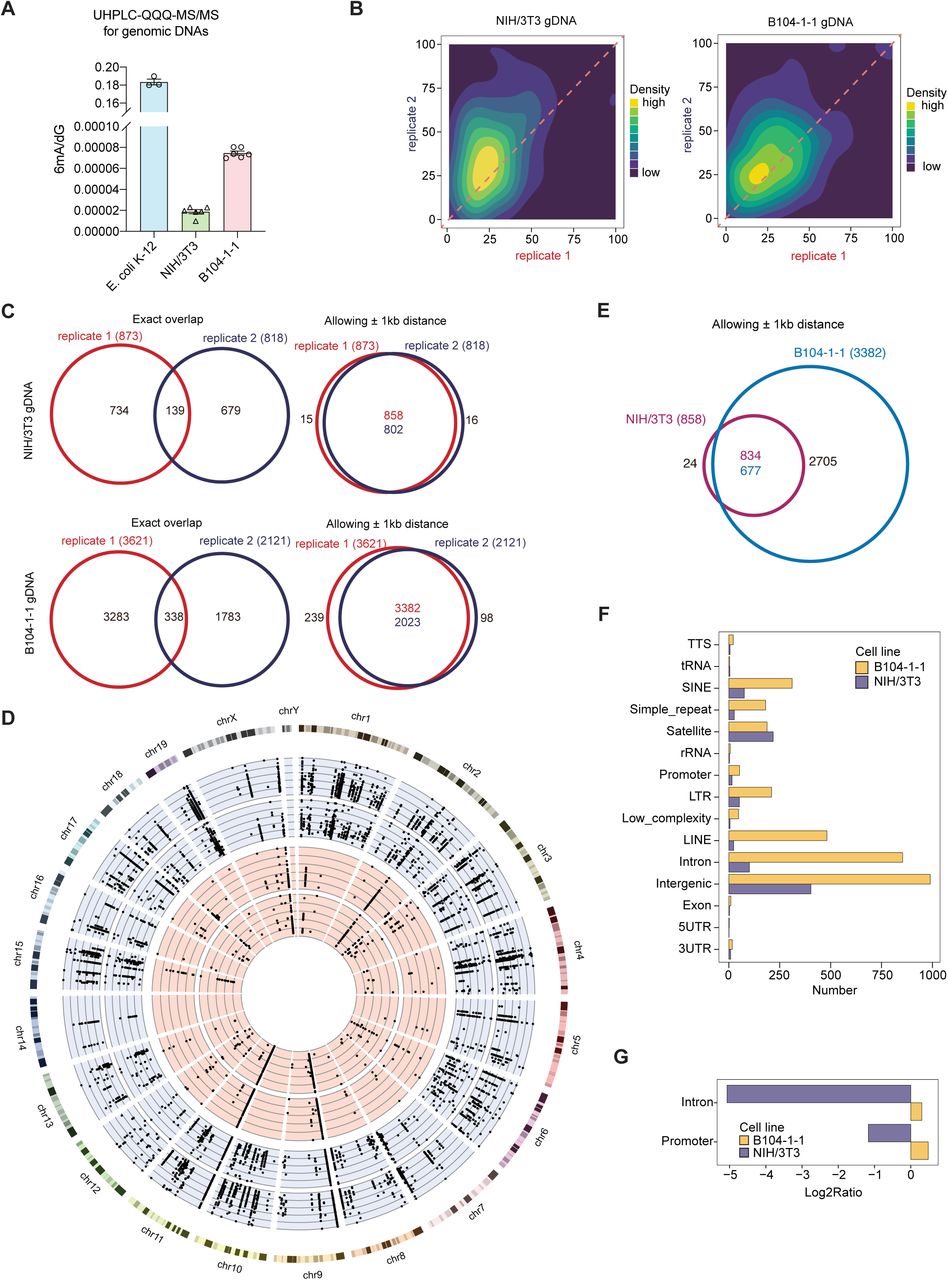
Figure Lengend Snippet: A) Quantification of 6mA level in gDNA from cultured NIH/3T3 and B104-1-1 cells, by LC-MS/MS, with E. coli gDNA as a positive control. Data are mean ± s.e.m. B) Correlation analysis of 6mA methylation fractions in antibody-enriched gDNA from NIH/3T3 and B104-1-1 cells, indicating a high correlation between replicates of DR-6mA-seq. C) Venn diagram showing the overlapped gDNA 6mA sites in NIH/3T3 and B104-1-1, allowing and not allowing ± 1 kb window, detected by DR-6mA-seq between two biologically independent replicates. D) Distributions of 6mA sites along the mouse genome. The dots on the blue and red background are gDNA 6mA sites in two biologically independent replicates of B104-1-1 and NIH/3T3 cells, respectively, revealed by DR-6mA-seq, with six circled axes representing the methylation fractions at 0%, 20%, 40%, 60%, 80%, and 100%. E) Venn diagram showing the overlapped gDNA 6mA sites between NIH/3T3 and B104-1-1 cells, allowing ± 1 kb window. F) Genomic distributions of gDNA 6mA sites in NIH/3T3 and B104-1-1 cells, revealed by DR-6mA-seq. G) Genomic enrichment of gDNA 6mA sites identified in NIH/3T3 and B104-1-1 cells, revealed by DR-6mA-seq. Enrichment scores on intron and promoter are shown. See also - and Tables S7-S8.
Article Snippet:
Techniques: Cell Culture, Liquid Chromatography with Mass Spectroscopy, Positive Control, Methylation
Journal: bioRxiv
Article Title: Sequencing of N 6 -methyl-deoxyadenosine at single-base resolution across the mammalian genome
doi: 10.1101/2023.01.16.524325
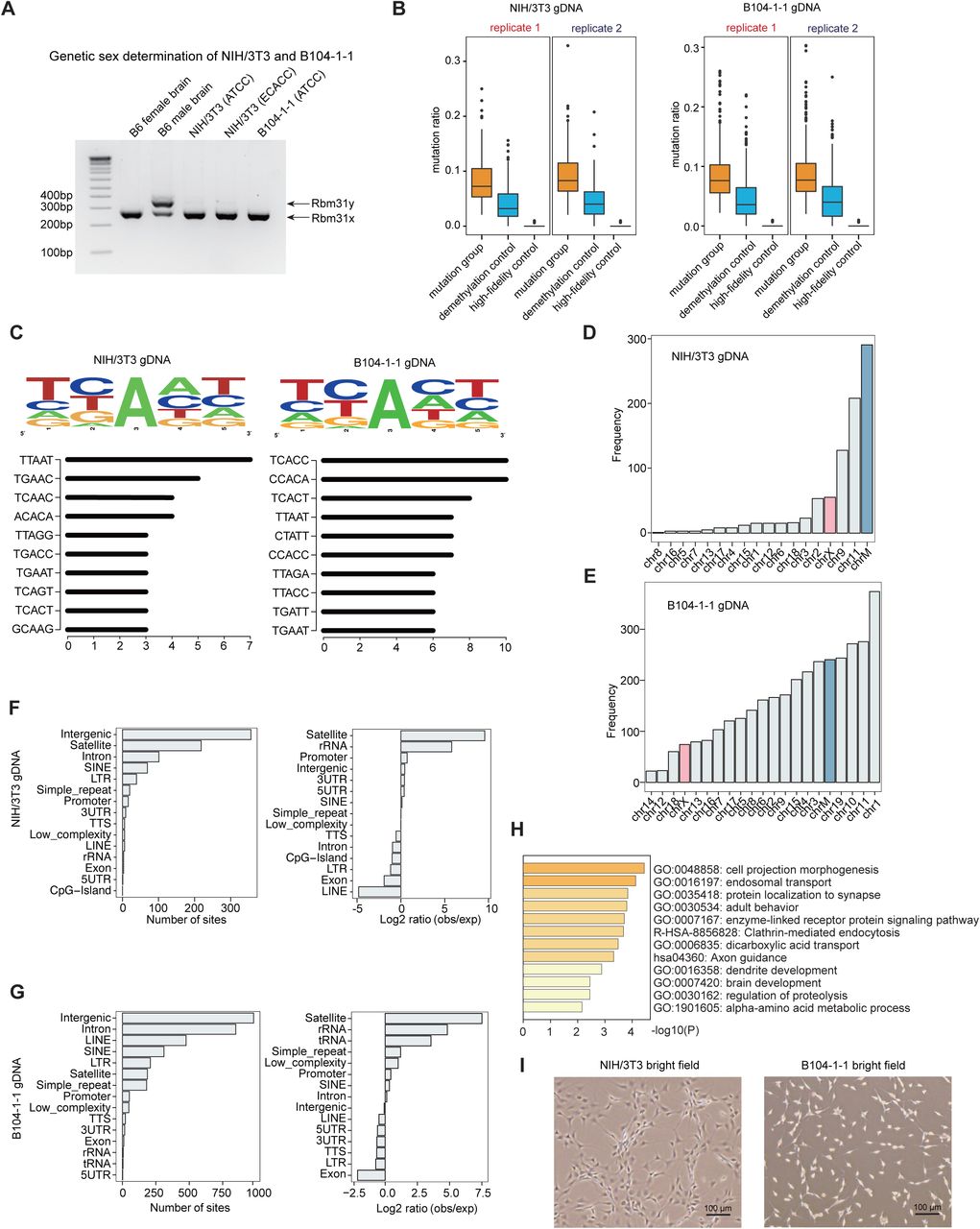
Figure Lengend Snippet: A) Genetic sex determination of NIH/3T3 and B104-1-1 cell lines by amplification of Rbm31x and Rbm31y by simplex PCR, with female and male mouse brains as controls. B) The box plot of mutation ratio distribution at 6mA sites detected in gDNA from NIH/3T3 (ATCC) and B104-1-1 (ATCC) cells, revealed by DR-6mA-seq. The mutation ratios of three groups are shown, such as Bst 2.0 extended DNA from untreated DNA (mutation group), Bst 2.0 extended DNA from FTO-treated DNA (demethylation control), and Q5-extended DNA from untreated DNA (high-fidelity control). C) Motif sequence logo and top 10 consensus motifs containing 6mA sites in gDNA from NIH/3T3 (ATCC) and B104-1-1 cells, uncovered by DR-6mA-seq. D) Chromosome-wide distributions of 6mA sites in gDNA from NIH/3T3. E) Chromosome-wide distributions of 6mA sites in gDNA from B104-1-1. F) Gene annotation shows the genomic distributions and enrichment of gDNA 6mA sites in NIH/3T3 cell lines, revealed by DR-6mA-seq. G) Gene annotation shows the genomic distributions and enrichment of gDNA 6mA sites identified in B104-1-1 cell lines, revealed by DR-6mA-seq. H) The enriched GO clusters of 6mA-modified genes in B104-1-1 cells. Genes with 6mA sites on exons, introns, and promoters are considered as 6mA-modified genes. I) Bright field photographs of NIH/3T3 and B104-1-1 cells.
Article Snippet:
Techniques: Amplification, Mutagenesis, Control, Sequencing, Modification
Journal: bioRxiv
Article Title: Sequencing of N 6 -methyl-deoxyadenosine at single-base resolution across the mammalian genome
doi: 10.1101/2023.01.16.524325
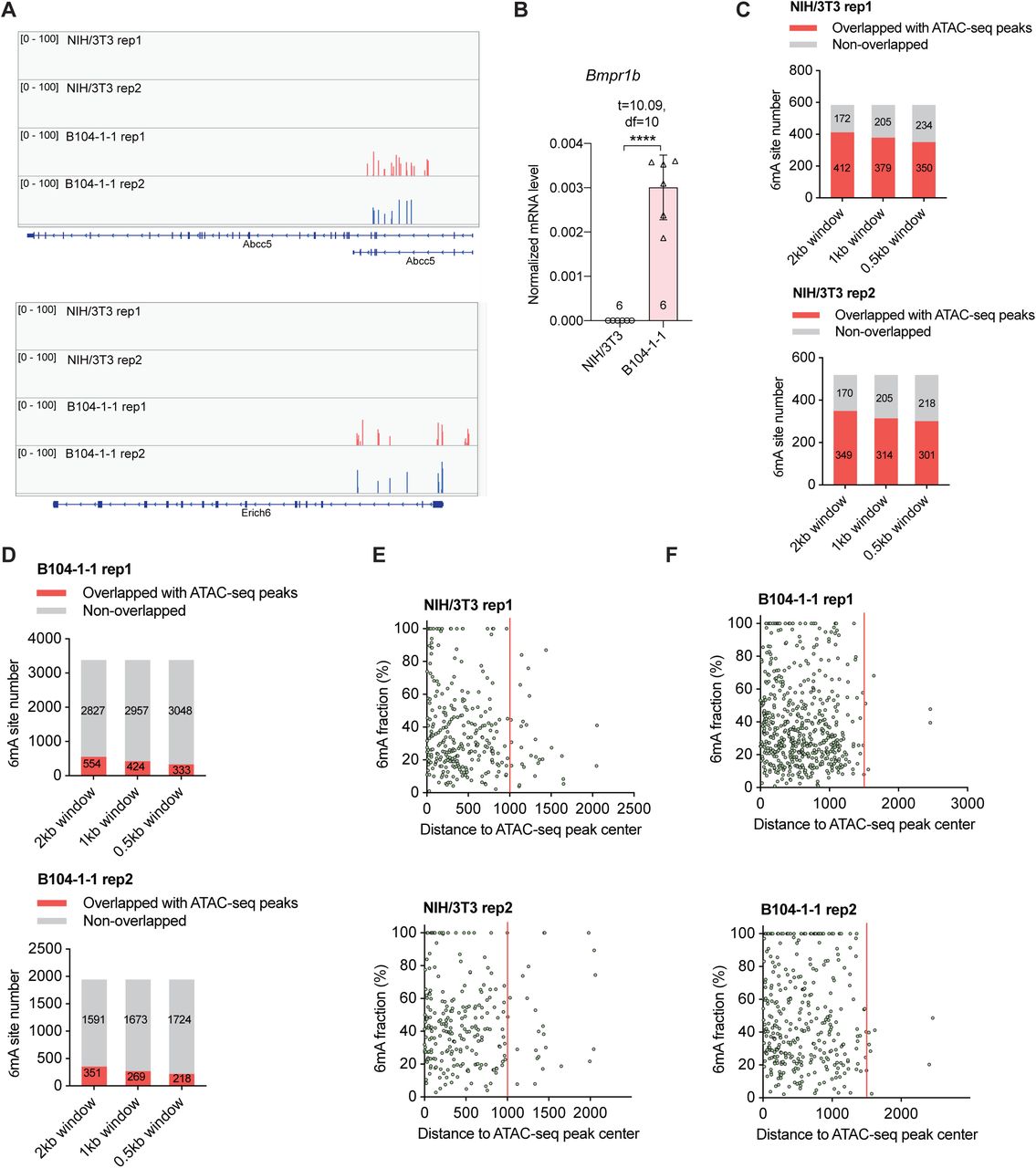
Figure Lengend Snippet: A) Two representative cell-line-specific gDNA 6mA clusters located at Abcc5 and Erich6 in B104-1-1 cells. B) RT-qPCR quantification of Bmpr1b expression level in NIH/3T3 and B104-1-1 cells, normalized to Actb . Data are mean ± s.e.m.; analyzed by two-tailed unpaired t-tests. The number of independently repeated reactions is shown in each plot. **** P <0.0001. C) The NIH/3T3 gDNA 6mA site number overlapped with ATAC-seq peaks from wild-type NIH/3T3 cells (GSE119781), by setting a sliding window of 500 bp, 1,000 bp, or 2,000 bp centered at the 6mA site revealed by DR-6mA-seq. D) The B104-1-1 gDNA 6mA site number overlapped with ATAC-seq peaks from glioblastoma tumor induced by implanted GL261 cells (GSE206551), by setting a sliding window of 500 bp, 1,000 bp, or 2,000 bp centered at the 6mA site revealed by DR-6mA-seq. E) The 2-D plot of NIH/3T3 gDNA 6mA methylation fraction versus the distance to ATAC-seq peak center (GSE119781), most 6mA sites distribute within ±1,000 bp range around ATAC-seq peak center. F) The 2-D plot of B104-1-1 gDNA 6mA methylation fraction versus the distance to ATAC-seq peak center (GSE206551), most 6mA sites distribute within ±1,500 bp range around ATAC-seq peak center.
Article Snippet:
Techniques: Quantitative RT-PCR, Expressing, Two Tailed Test, Methylation
Journal: bioRxiv
Article Title: Sequencing of N 6 -methyl-deoxyadenosine at single-base resolution across the mammalian genome
doi: 10.1101/2023.01.16.524325
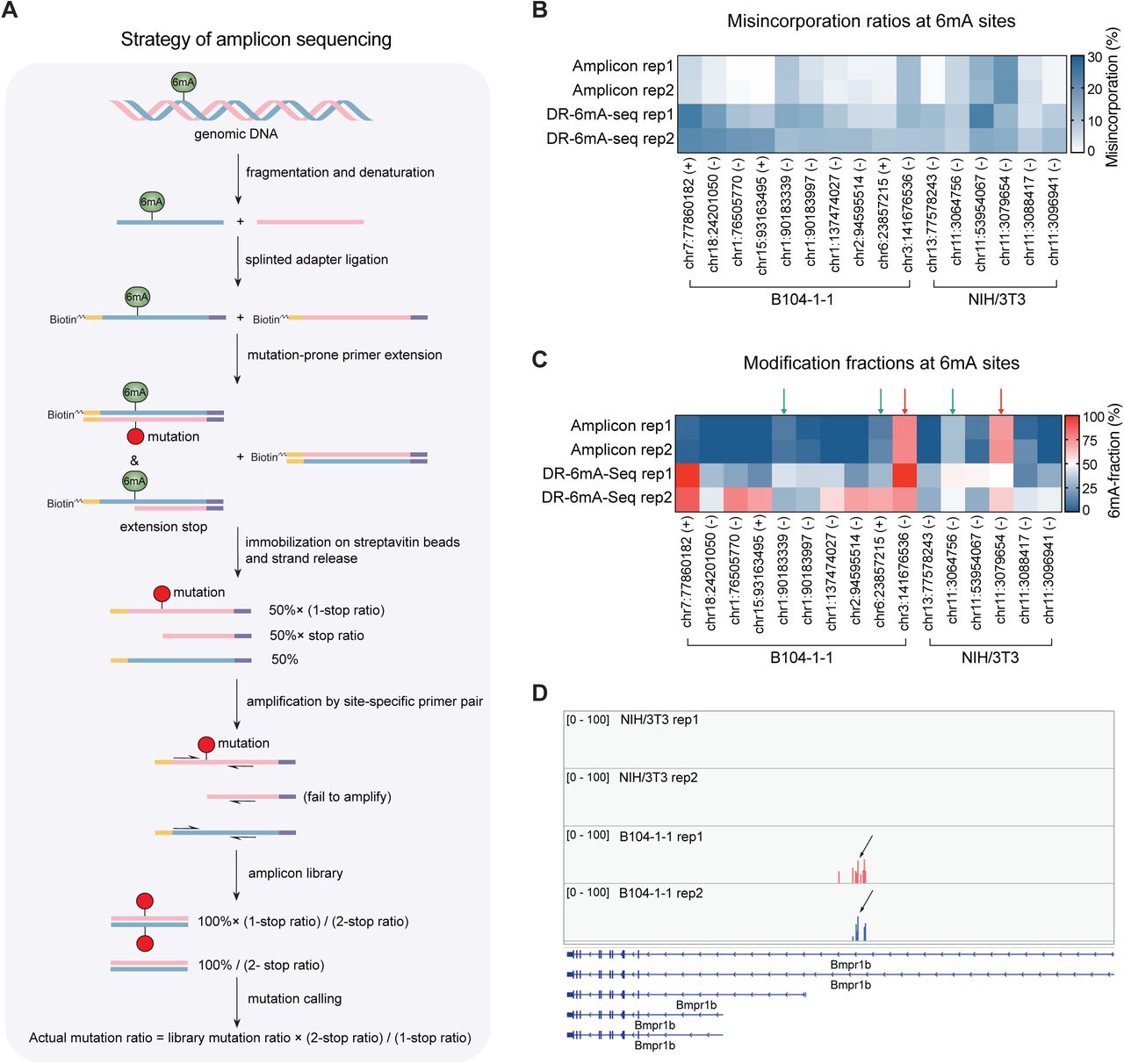
Figure Lengend Snippet: A) A flowchart of amplicon assay for gDNA 6mA validation, revealing 6mA fraction by adjusted misincorporation ratios. B) The actual misincorporation ratios obtained in amplicon assay versus the misincorporation calculated from DR-6mA-seq data. C) The 6mA methylation fractions by amplicon assay versus 6mA fractions calculated from DR-6mA-seq data. gDNA 6mA sites showing >70% estimated modification in amplicon assay were marked by red arrows; other gDNA 6mA sites showing above 10% modification fraction were labeled by green arrows. For B) and C), 16 gDNA 6mA sites were investigated, with 6 from NIH/3T3 cells and 10 from B104-1-1 cells. D) A representative 6mA cluster located at Bmpr1b , with the arrow marking the 6mA site of the highest modification fraction identified in C). See also - .
Article Snippet:
Techniques: Amplification, Biomarker Discovery, Methylation, Modification, Labeling